What: the Glycine Receptor Allosteric Ligand Library (GRALL) is the first database of allosteric modulators of a human synaptic receptor with a structural annotation based on their binding site on the receptor. GRALL offers a collection of 218 chemical entities with documented modulatory activities at homomeric glycine receptors (GlyR) α1 and α3, which includes agonists, antagonists, positive and negative allosteric modulators and large number of experimentally inactive compounds. The GRALL database provides an extensive overview of the pharmacology of a relevant drug target that is expected to boost the development of in silico methodologies for allosteric drug design.
Why: GRALL is a unique contribution to the field of neuropharmacology. This collection provides information on the chemical structure, the direction of modulation, the potency, the 3D molecular structure and quantum-mechanical charges of a large number of biologically active compounds at a human synaptic receptor. Most importantly, a large fraction of modulators in GRALL comes with a structural annotation of their ligand-binding site on the receptor, which was assigned using a level of confidence from 1 (highest) to 5 (lowest) depending on the quality of the evidence supporting the annotation. This information, which is currently missing in popular drug banks like ChEMBL, PubChem, or Binding DB provides a stringent benchmark that is expected to boost the development of predictive in silico strategies for allosteric drug design. Moreover, it can be used to combine ligand-based and structure-based approaches for drug design.
How: GRALL is organized as a web-accessible database, where compounds can be ranked by name, SMILES, chemical family, annotated binding site, level of confidence of annotation, the direction of modulation, the activity type and value. To extract information, the database can be searched using keywords (see below), or downloaded in the CSV format. For each entry, the database allows for immediate visualization of the 2D chemical structure and provides the DOI of the most relevant publication associated to it. The hyperlink in the last column provides access to the 3D representation in MOL2 format, which is suitable for straightforward visualization. The whole chemical library can also be directly downloaded in a multi MOL2 format for direct docking or virtual screening.
Contact: Dr. Marco Cecchini (mcecchini@unistra.fr)
Reference: Cerdan A.H., Sisquellas M., Pereira G., Gomes D.E.B., Changeux J-P. & Cecchini M. “The glycine receptor allosteric ligands library (GRALL)”. Bioinformatics 36, 3379–3384 (2020). https://doi.org/10.1093/bioinformatics/btaa170
Annotation:
- Chemical family: alcohol, alkaloid, aminoacid, avermectin, bilabolide, cannabinoid, cyanotriphenylborate, dihydropyridine, general_anesthetic, ginkgolic-acid, ginkgolide, glucocortico_steroid, glutamate, HTS, neurosteroid, phenylalanine, phenylpyrazole, picrotoxin, propofol, sulfonamide, tropeine.
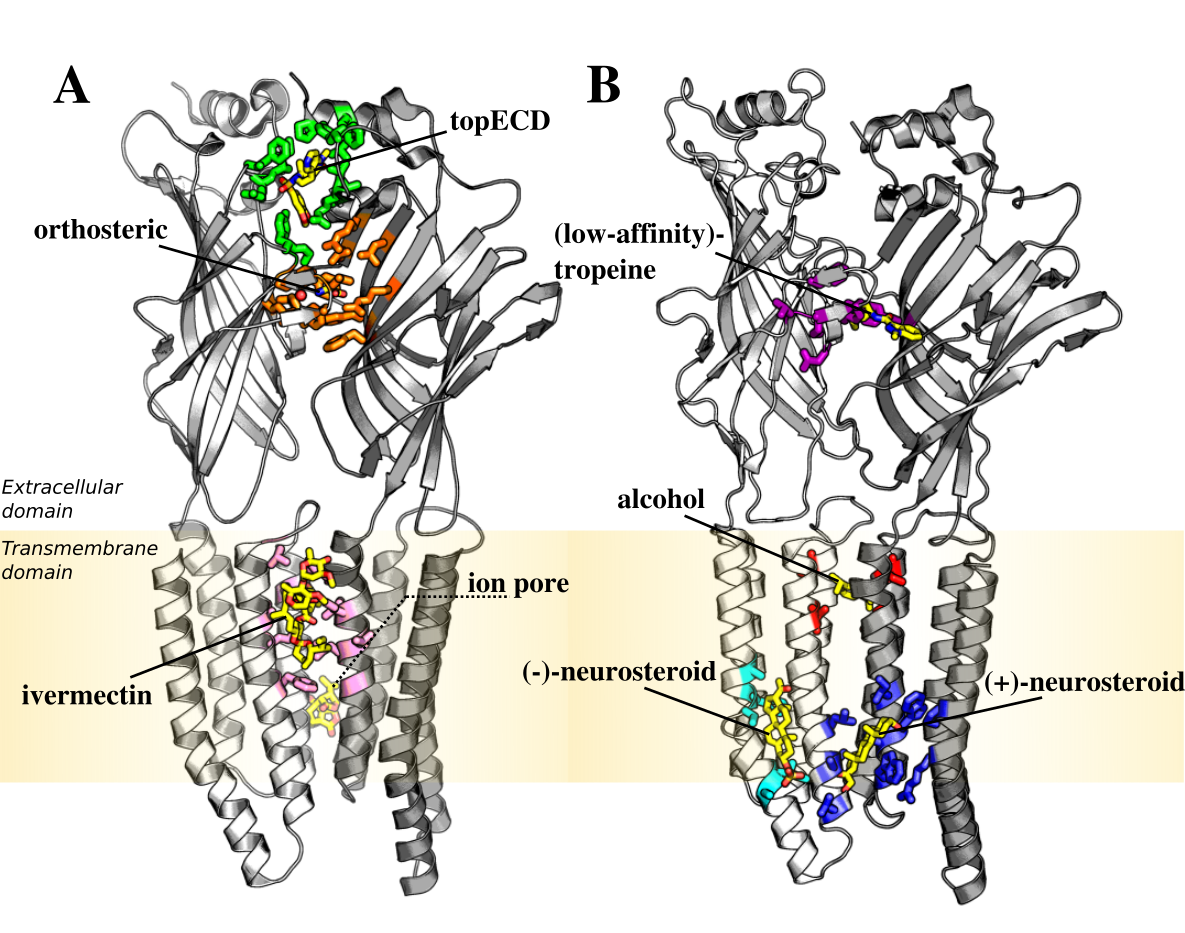
- Binding site: (+)-neurosteroid, (−)-neurosteroid, (LA)-tropeine, alcohol, ivermectin, orthosteric, pore, top_ECD or N.A.
- Effect:
0: Inactive
+: Agonism or positive modulation
−: Antagonism or negative modulation
N.A.: Not Available - Activity type: EC50, IC50, Kd, Maximum_effect(%),Mean_potentiation(%) or N.A.
- Level of confidence:
1: High-Resolution Structure from GlyR
2: High-Resolution Structure from other pLGIC + concordant evidence in GlyR
3: Site-Directed Mutagenesis
4: Modelling / Computational Studies
5: Structural Similarities with annotated ligands
N.A: Not Available
Update: We encourage researchers to contact us and submit missing or new molecules via the preformatted form (Submit new ligand). Upon verification, the transmitted information will be added to the database and published online.
| Name | Structure | SMILES | Family | Binding Site | Level of Confidence | Effect α1 | Effect α3 | Activity Type α1 | Activity Value α1 | Activity Type α3 | Activity Value α3 | DOI | Download |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| conivaptan | 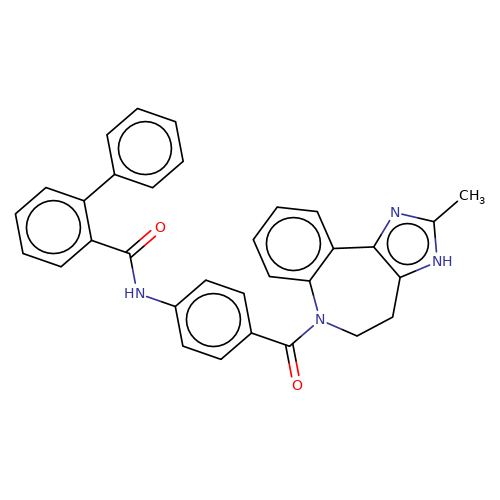 | Cc1nc2c([nH]1)CCN(C(=O)c1ccc(NC(=O)c3ccccc3-c3ccccc3)cc1)c1ccccc1-2 | HTS | N.A | N.A | 0 | N.A | EC50(uM) | Inactive | N.A | N.A | 10.1021/jm501873p | |
| PNU-120596 | 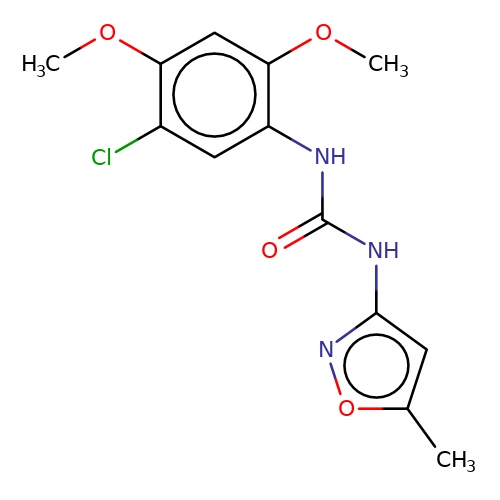 | COc1cc(OC)c(NC(=O)Nc2cc(C)on2)cc1Cl | HTS | N.A | N.A | 0 | N.A | EC50(uM) | Inactive | N.A | N.A | 10.1021/jm501873p | |
| pimozide | 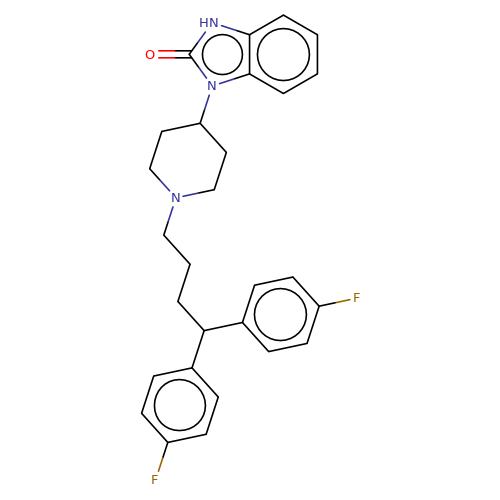 | O=c1[nH]c2ccccc2n1C1CCN(CCCC(c2ccc(F)cc2)c2ccc(F)cc2)CC1 | HTS | N.A | N.A | + | N.A | EC50(uM) | 1.7 | N.A | N.A | 10.1021/jm501873p | |
| cholecalciferol | 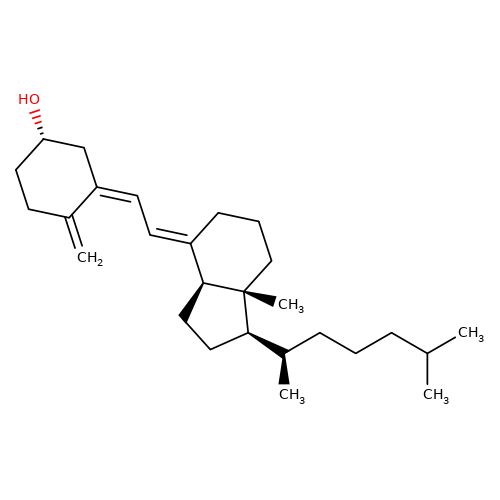 | C=C1CC[C@H](O)C/C1=C/C=C1\CCC[C@@]2(C)[C@H]1CC[C@@H]2[C@H](C)CCCC(C)C | HTS | N.A | N.A | + | N.A | EC50(uM) | 0.4 | N.A | N.A | 10.1021/jm501873p | |
| cinacalcet | 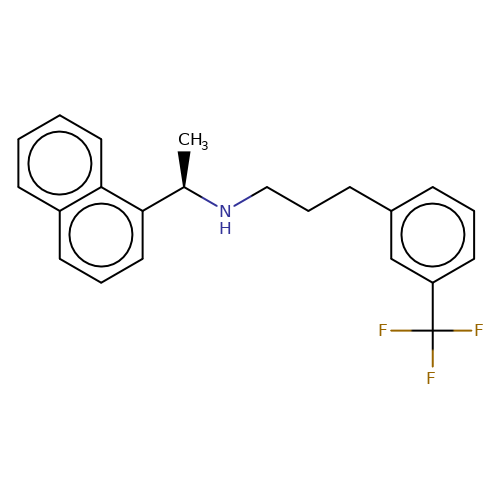 | C[C@@H](NCCCc1cccc(C(F)(F)F)c1)c1cccc2ccccc12 | HTS | N.A | N.A | + | N.A | EC50(uM) | 0.32 | N.A | N.A | 10.1021/jm501873p | |
| dutasteride | 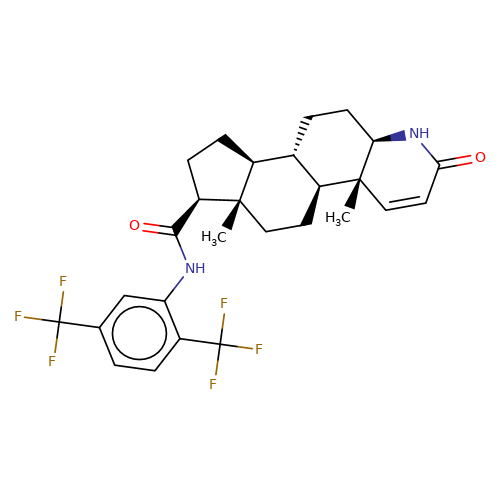 | C[C@]12CC[C@H]3[C@@H](CC[C@H]4NC(=O)C=C[C@]34C)[C@@H]1CC[C@@H]2C(=O)Nc1cc(C(F)(F)F)ccc1C(F)(F)F | HTS | N.A | N.A | + | N.A | EC50(uM) | 0.33 | N.A | N.A | 10.1021/jm501873p | |
| sulindac | 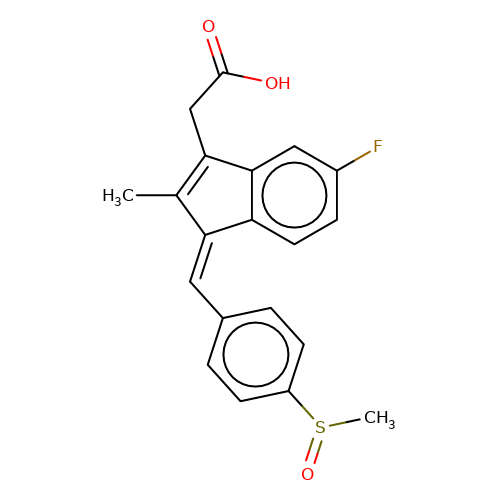 | CC1=C(CC(=O)O)c2cc(F)ccc2/C1=C\c1ccc(S(C)=O)cc1 | HTS | N.A | N.A | + | N.A | EC50(uM) | 0.38 | N.A | N.A | 10.1021/jm501873p | |
| risperidone | 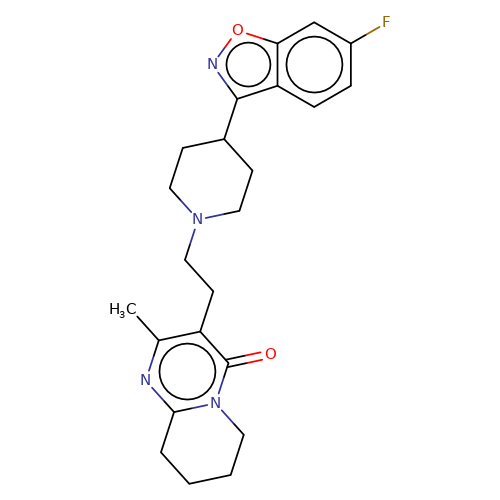 | Cc1nc2n(c(=O)c1CCN1CCC(c3noc4cc(F)ccc34)CC1)CCCC2 | HTS | N.A | N.A | + | N.A | EC50(uM) | 0.32 | N.A | N.A | 10.1021/jm501873p | |
| adapalene | 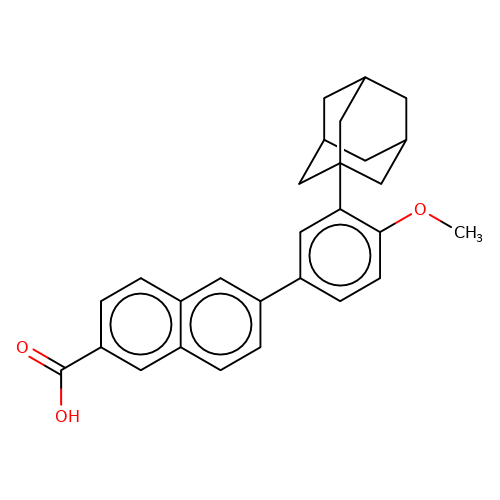 | COc1ccc(-c2ccc3cc(C(=O)O)ccc3c2)cc1C12CC3CC(CC(C3)C1)C2 | HTS | N.A | N.A | + | N.A | EC50(uM) | 1.3 | N.A | N.A | 10.1021/jm501873p | |
| fluspirilene | 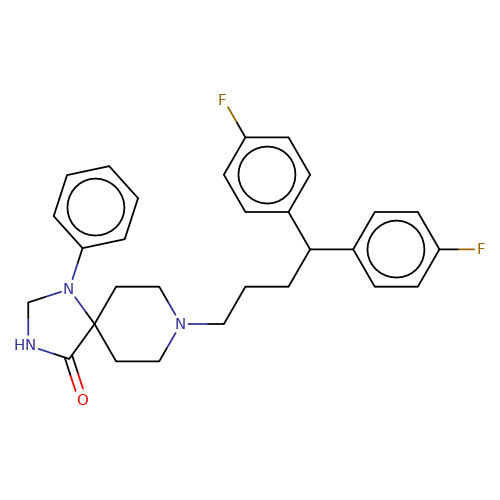 | O=C1NCN(c2ccccc2)C12CCN(CCCC(c1ccc(F)cc1)c1ccc(F)cc1)CC2 | HTS | N.A | N.A | + | N.A | EC50(uM) | 1.2 | N.A | N.A | 10.1021/jm501873p | |
| mefloquine | 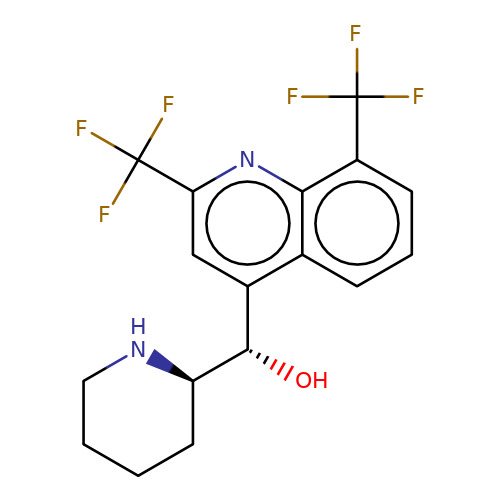 | O[C@@H](c1cc(C(F)(F)F)nc2c(C(F)(F)F)cccc12)[C@H]1CCCCN1 | HTS | N.A | N.A | + | N.A | EC50(uM) | 2.5 | N.A | N.A | 10.1021/jm501873p | |
| astemizole | 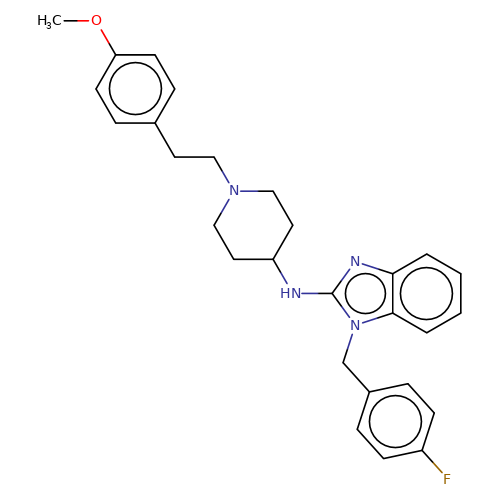 | COc1ccc(CCN2CCC(Nc3nc4ccccc4n3Cc3ccc(F)cc3)CC2)cc1 | HTS | N.A | N.A | + | N.A | EC50(uM) | 4.2 | N.A | N.A | 10.1021/jm501873p | |
| telmisartan | 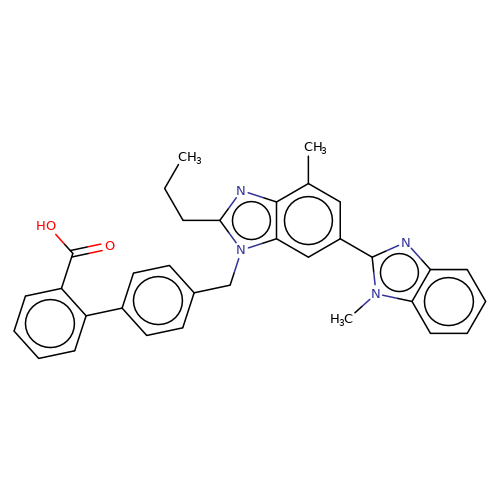 | CCCc1nc2c(C)cc(-c3nc4ccccc4n3C)cc2n1Cc1ccc(-c2ccccc2C(=O)O)cc1 | HTS | N.A | N.A | + | N.A | EC50(uM) | 3.2 | N.A | N.A | 10.1021/jm501873p | |
| regorafenib | 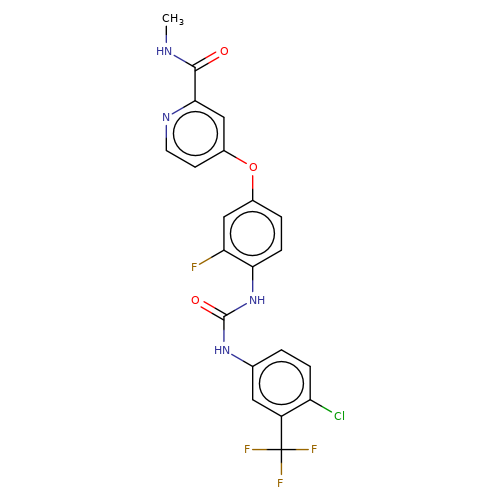 | CNC(=O)c1cc(Oc2ccc(NC(=O)Nc3ccc(Cl)c(C(F)(F)F)c3)c(F)c2)ccn1 | HTS | N.A | N.A | + | N.A | EC50(uM) | 1.8 | N.A | N.A | 10.1021/jm501873p | |
| balansa-a-1 | 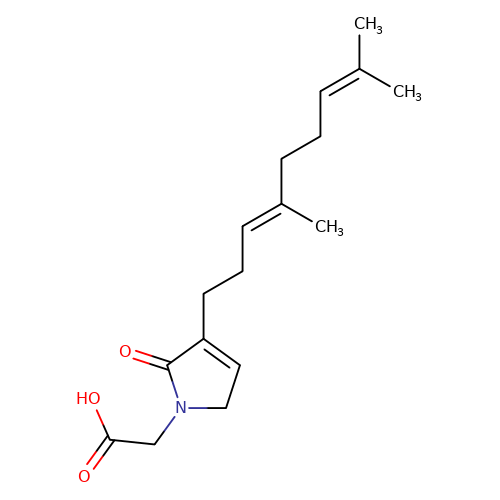 | CC(C)=CCC/C(C)=C/CCC1=CCN(CC(=O)O)C1=O | HTS | N.A | N.A | 0 | 0 | IC50(uM) | Inactive | IC50(uM) | Inactive | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-2 | 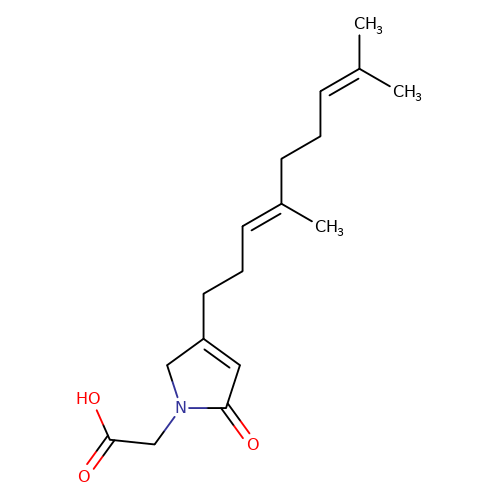 | CC(C)=CCC/C(C)=C/CCC1=CC(=O)N(CC(=O)O)C1 | HTS | N.A | N.A | 0 | 0 | IC50(uM) | Inactive | IC50(uM) | Inactive | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-3 | 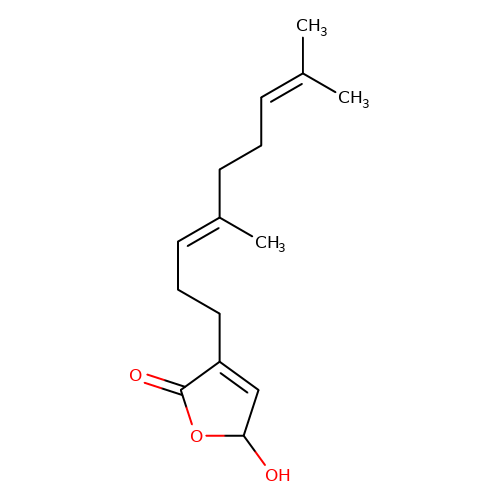 | CC(C)=CCC/C(C)=C/CCC1=CC(O)OC1=O | HTS | N.A | N.A | 0 | 0 | IC50(uM) | Inactive | IC50(uM) | Inactive | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-4 | 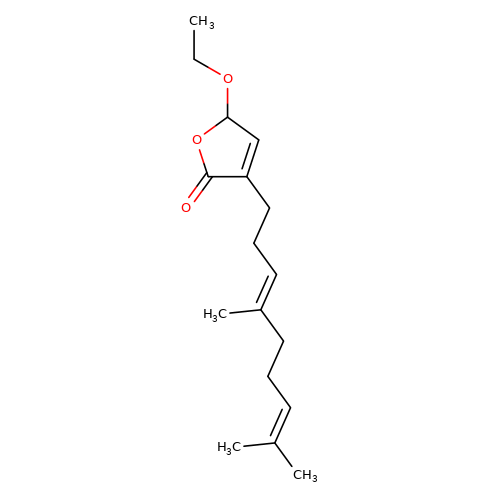 | CCOC1C=C(CC/C=C(\C)CCC=C(C)C)C(=O)O1 | HTS | N.A | N.A | 0 | 0 | IC50(uM) | Inactive | IC50(uM) | Inactive | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-5 | 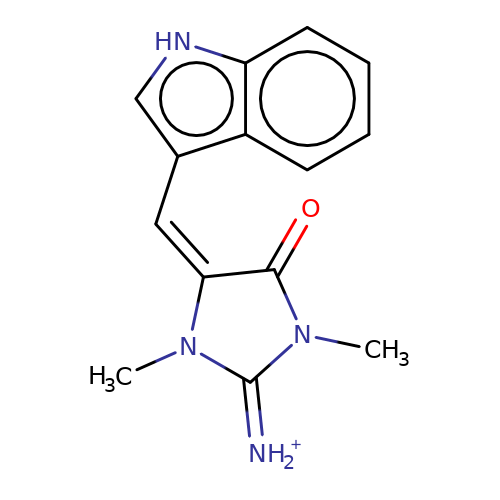 | CN1C(=[NH2+])N(C)/C(=C/c2c[nH]c3ccccc23)C1=O | HTS | N.A | N.A | 0 | 0 | IC50(uM) | Inactive | IC50(uM) | Inactive | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-8 | 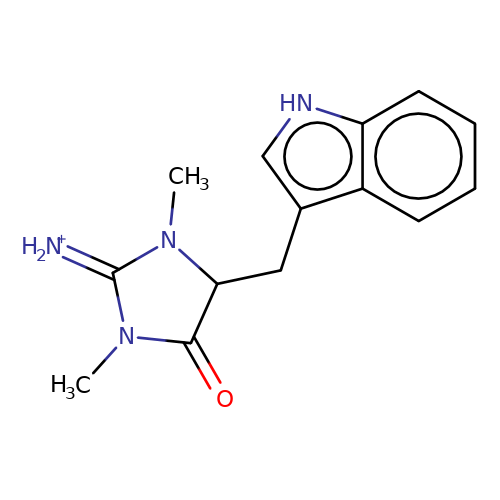 | CN1C(=[NH2+])N(C)C(Cc2c[nH]c3ccccc23)C1=O | HTS | N.A | N.A | 0 | 0 | IC50(uM) | Inactive | IC50(uM) | Inactive | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-10 | 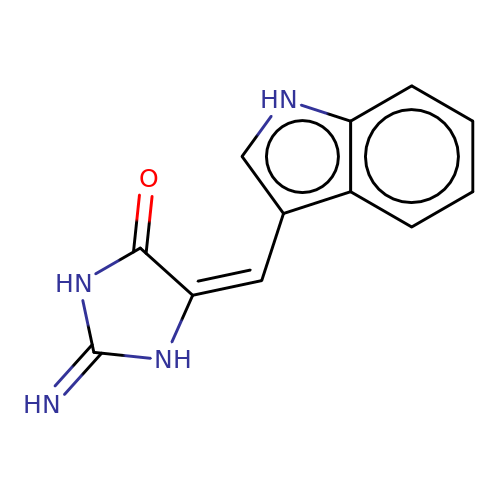 | N=C1NC(=O)/C(=C\c2c[nH]c3ccccc23)N1 | HTS | N.A | N.A | 0 | 0 | IC50(uM) | Inactive | IC50(uM) | Inactive | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-12a | 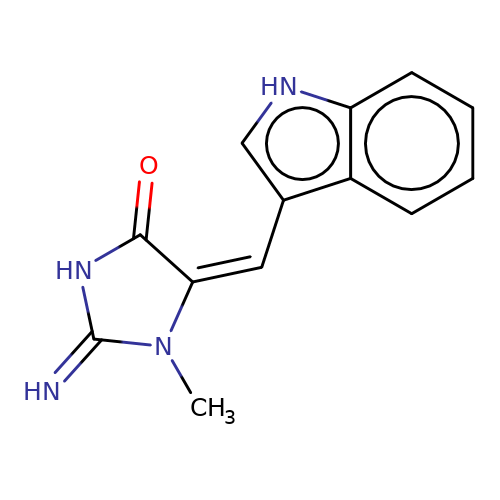 | CN1C(=N)NC(=O)/C1=C\c1c[nH]c2ccccc12 | HTS | N.A | N.A | 0 | 0 | IC50(uM) | Inactive | IC50(uM) | Inactive | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-12b | 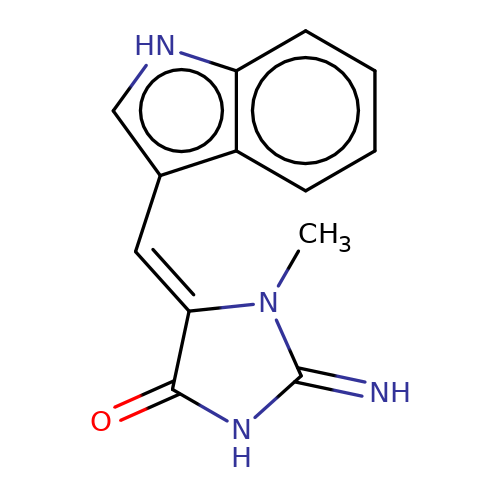 | CN1C(=N)NC(=O)/C1=C/c1c[nH]c2ccccc12 | HTS | N.A | N.A | 0 | 0 | IC50(uM) | Inactive | IC50(uM) | Inactive | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-13 | 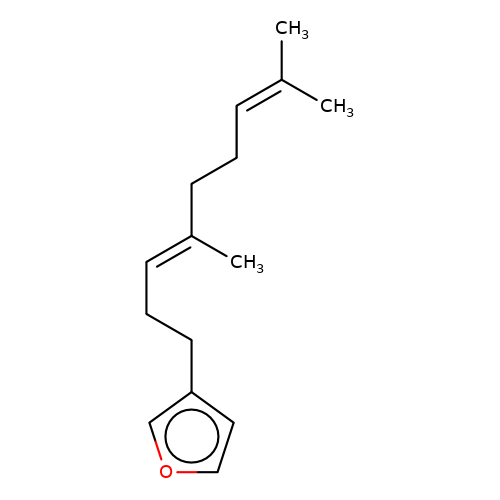 | CC(C)=CCC/C(C)=C/CCc1ccoc1 | HTS | N.A | N.A | 0 | 0 | IC50(uM) | Inactive | IC50(uM) | Inactive | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-6 | 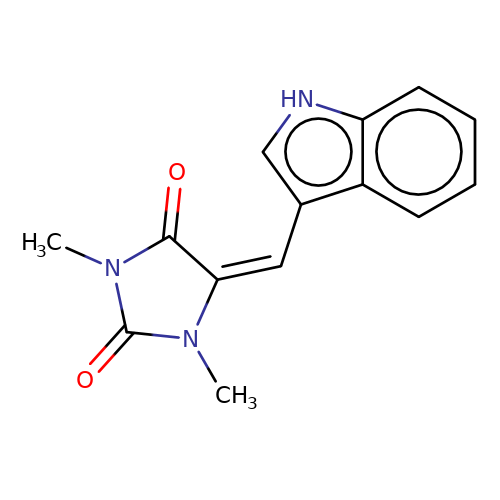 | CN1C(=O)/C(=C\c2c[nH]c3ccccc23)N(C)C1=O | HTS | N.A | N.A | - | - | IC50(uM) | 200 | IC50(uM) | 67 | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-7 | 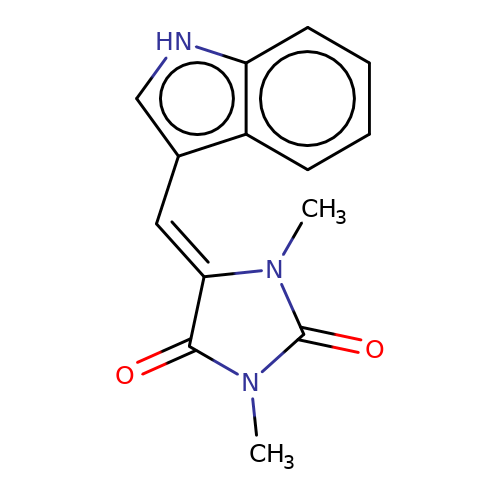 | CN1C(=O)/C(=C/c2c[nH]c3ccccc23)N(C)C1=O | HTS | N.A | N.A | - | - | IC50(uM) | 200 | IC50(uM) | 67 | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-9 | 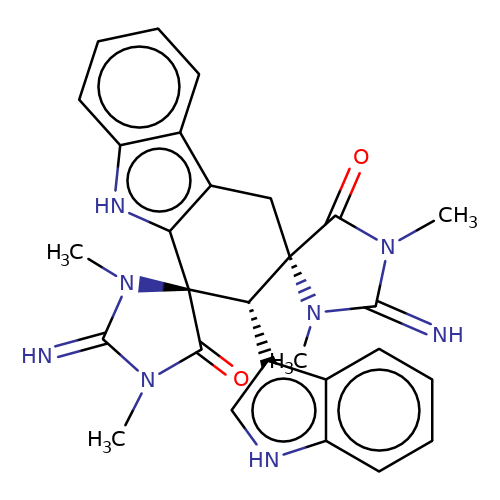 | CN1C(=N)N(C)[C@]2(Cc3c([nH]c4ccccc34)[C@@]3(C(=O)N(C)C(=N)N3C)[C@H]2c2c[nH]c3ccccc23)C1=O | HTS | N.A | N.A | - | - | IC50(uM) | 27 | IC50(uM) | 300 | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-11a | 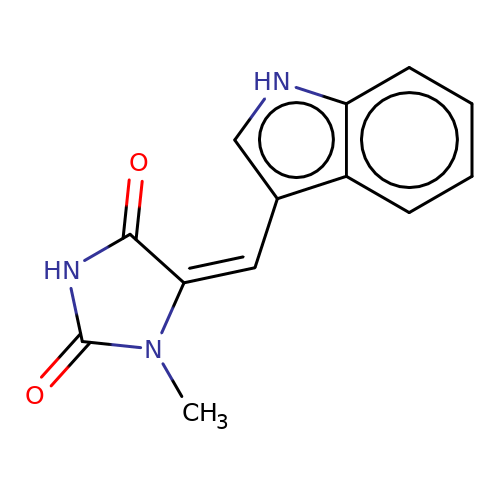 | CN1C(=O)NC(=O)/C1=C\c1c[nH]c2ccccc12 | HTS | N.A | N.A | - | - | IC50(uM) | 9 | IC50(uM) | 34 | 10.1016/j.bmc.2013.04.061 | |
| balansa-a-11b | 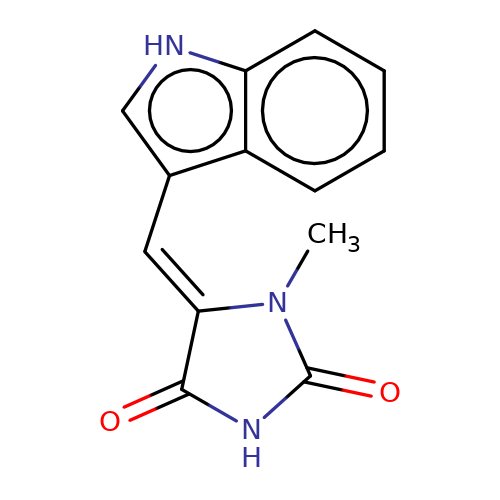 | CN1C(=O)NC(=O)/C1=C/c1c[nH]c2ccccc12 | HTS | N.A | N.A | - | - | IC50(uM) | 9 | IC50(uM) | 34 | 10.1016/j.bmc.2013.04.061 | |
| balansa-b-2 | 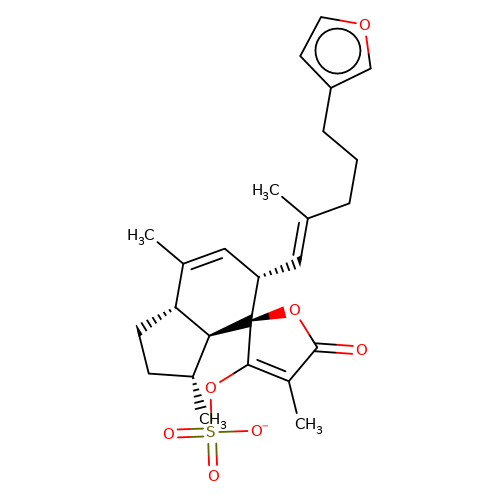 | CC1=C[C@H](/C=C(\C)CCCc2ccoc2)[C@@]2(OC(=O)C(C)=C2OS(=O)(=O)[O-])[C@@H]2[C@@H]1CC[C@H]2C | HTS | N.A | N.A | - | - | Maximum_effect(%) | 34 | Maximum_effect(%) | 17 | 10.1039/c3ob40861b | |
| balansa-b-1 | 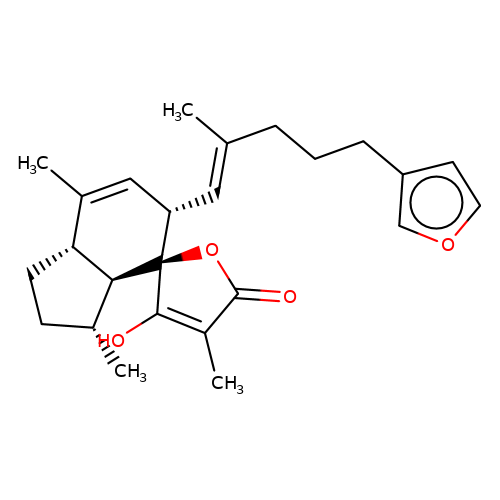 | CC1=C[C@H](/C=C(\C)CCCc2ccoc2)[C@@]2(OC(=O)C(C)=C2O)[C@@H]2[C@@H]1CC[C@H]2C | HTS | N.A | N.A | + | - | Maximum_effect(%) | 175 | Maximum_effect(%) | 41 | 10.1039/c3ob40861b | |
| balansa-b-5 | 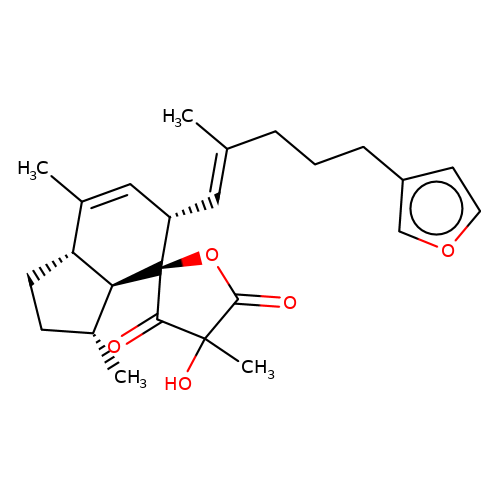 | CC1=C[C@H](/C=C(\C)CCCc2ccoc2)[C@@]2(OC(=O)C(C)(O)C2=O)[C@@H]2[C@@H]1CC[C@H]2C | HTS | N.A | N.A | + | - | Maximum_effect(%) | 115 | Maximum_effect(%) | 48 | 10.1039/c3ob40861b | |
| balansa-b-6 | 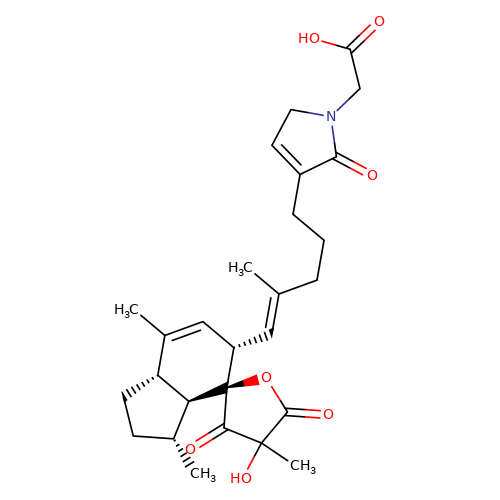 | CC1=C[C@H](/C=C(\C)CCCC2=CCN(CC(=O)O)C2=O)[C@@]2(OC(=O)C(C)(O)C2=O)[C@@H]2[C@@H]1CC[C@H]2C | HTS | N.A | N.A | + | - | Maximum_effect(%) | 211 | Maximum_effect(%) | 57 | 10.1039/c3ob40861b | |
| balansa-b-7 | 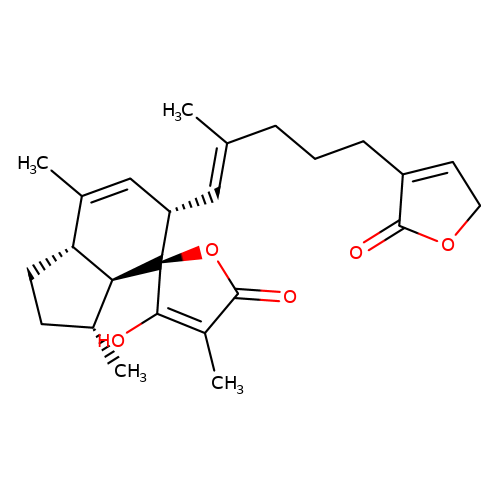 | CC1=C[C@H](/C=C(\C)CCCC2=CCOC2=O)[C@@]2(OC(=O)C(C)=C2O)[C@@H]2[C@@H]1CC[C@H]2C | HTS | N.A | N.A | + | - | Maximum_effect(%) | 118 | Maximum_effect(%) | 84 | 10.1039/c3ob40861b | |
| balansa-b-8 | 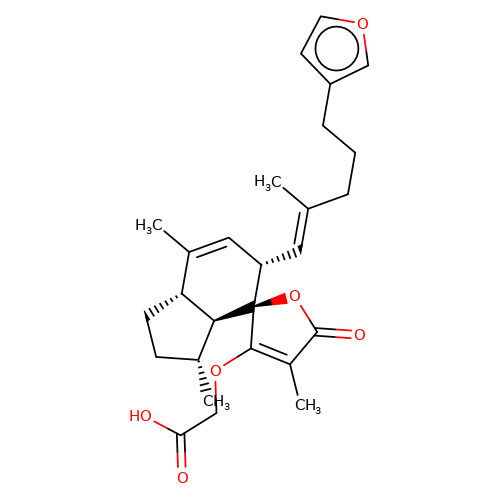 | CC1=C[C@H](/C=C(\C)CCCc2ccoc2)[C@@]2(OC(=O)C(C)=C2OCC(=O)O)[C@@H]2[C@@H]1CC[C@H]2C | HTS | N.A | N.A | + | - | Maximum_effect(%) | 121 | Maximum_effect(%) | 74 | 10.1039/c3ob40861b | |
| stead-1 | 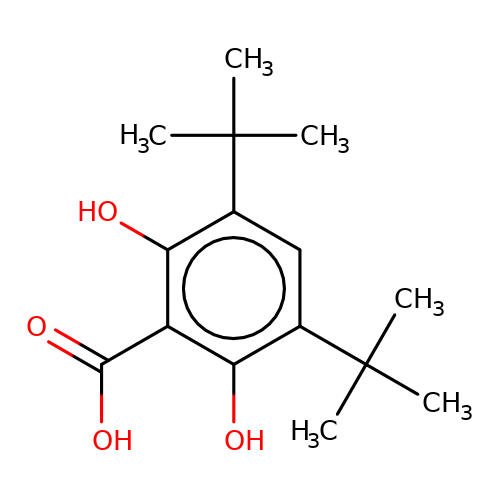 | CC(C)(C)c1cc(C(C)(C)C)c(O)c(C(=O)O)c1O | HTS | N.A | N.A | N.A | + | N.A | N.A | Mean_potentiation(%) | 2061 | 10.1177/1087057116657779 | |
| stead-2 | 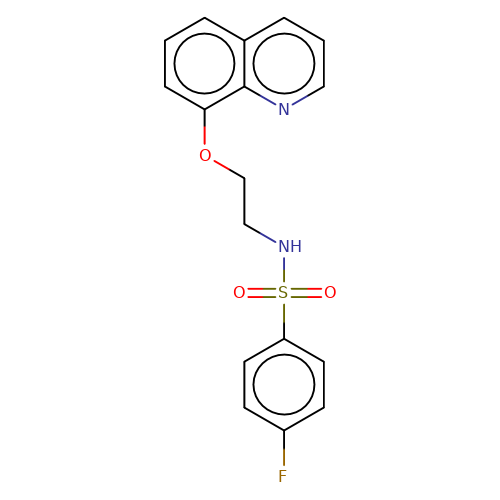 | O=S(=O)(NCCOc1cccc2cccnc12)c1ccc(F)cc1 | HTS | N.A | N.A | N.A | + | N.A | N.A | Mean_potentiation(%) | 2710 | 10.1177/1087057116657779 | |
| stead-3 | 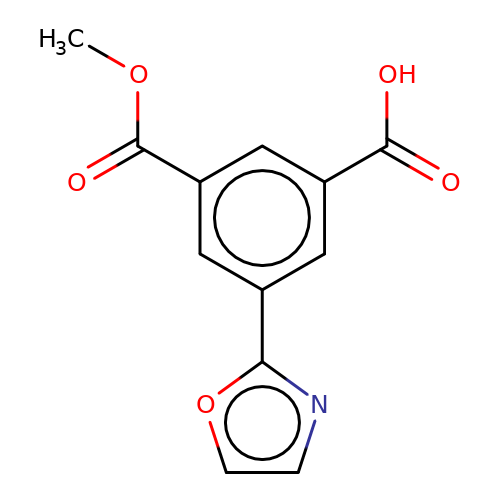 | COC(=O)c1cc(C(=O)O)cc(-c2ncco2)c1 | HTS | N.A | N.A | N.A | + | N.A | N.A | Mean_potentiation(%) | 255 | 10.1177/1087057116657779 | |
| stead-4 | 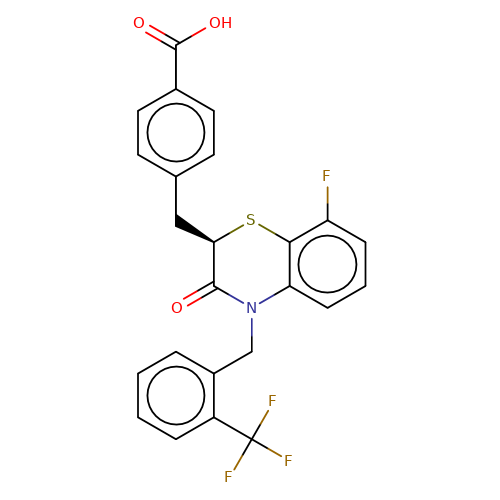 | O=C(O)c1ccc(C[C@H]2Sc3c(F)cccc3N(Cc3ccccc3C(F)(F)F)C2=O)cc1 | HTS | N.A | N.A | N.A | + | N.A | N.A | Mean_potentiation(%) | 396 | 10.1177/1087057116657779 | |
| stead-5 | 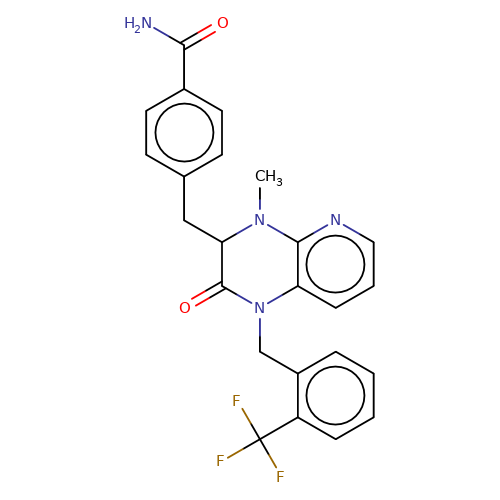 | CN1c2ncccc2N(Cc2ccccc2C(F)(F)F)C(=O)C1Cc1ccc(C(N)=O)cc1 | HTS | N.A | N.A | N.A | + | N.A | N.A | Mean_potentiation(%) | 303 | 10.1177/1087057116657779 | |
| stead-6 | 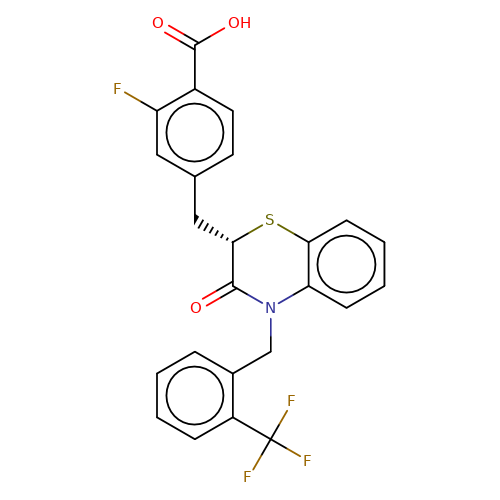 | O=C(O)c1ccc(C[C@@H]2Sc3ccccc3N(Cc3ccccc3C(F)(F)F)C2=O)cc1F | HTS | N.A | N.A | N.A | + | N.A | N.A | Mean_potentiation(%) | 259 | 10.1177/1087057116657779 | |
| stead-7 | 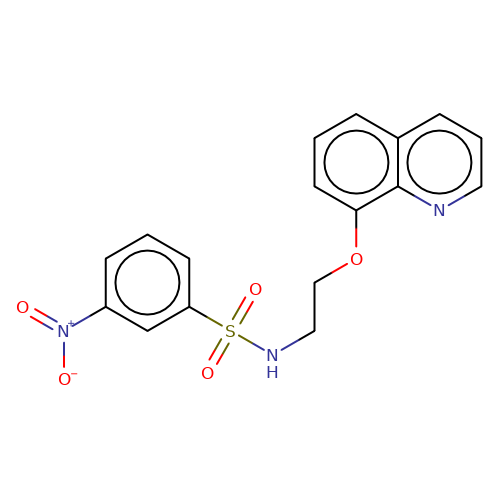 | O=[N+]([O-])c1cccc(S(=O)(=O)NCCOc2cccc3cccnc23)c1 | HTS | N.A | N.A | N.A | + | N.A | N.A | Mean_potentiation(%) | 136 | 10.1177/1087057116657779 | |
| balansa-b-3 | 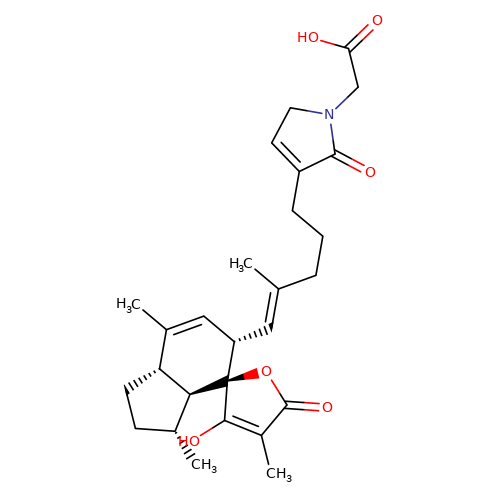 | CC1=C[C@H](/C=C(\C)CCCC2=CCN(CC(=O)O)C2=O)[C@@]2(OC(=O)C(C)=C2O)[C@@H]2[C@@H]1CC[C@H]2C | HTS | N.A | N.A | + | + | Maximum_effect(%) | 123 | Maximum_effect(%) | 367 | 10.1039/c3ob40861b | |
| balansa-b-4 | 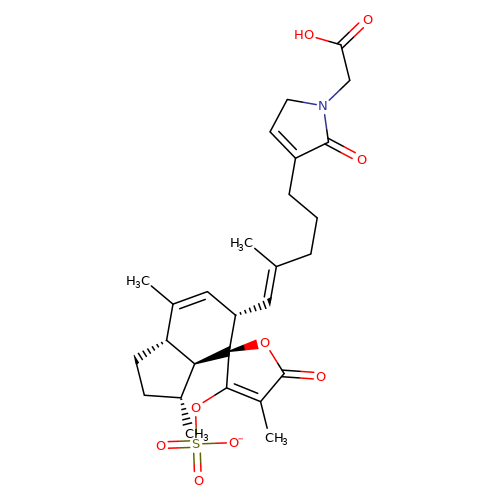 | CC1=C[C@H](/C=C(\C)CCCC2=CCN(CC(=O)O)C2=O)[C@@]2(OC(=O)C(C)=C2OS(=O)(=O)[O-])[C@@H]2[C@@H]1CC[C@H]2C | HTS | N.A | N.A | + | + | Maximum_effect(%) | 190 | Maximum_effect(%) | 190 | 10.1039/c3ob40861b | |
| ethanol | 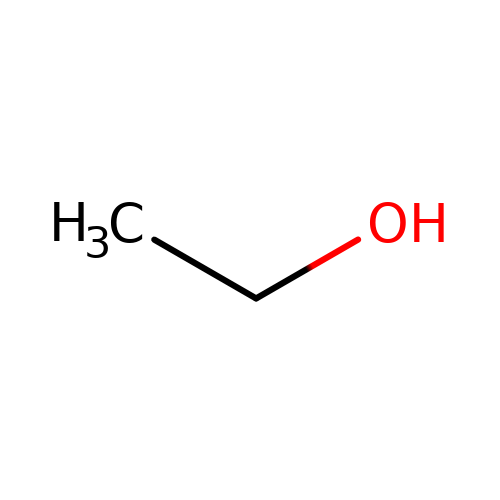 | CCO | alcohol | alcohol | 2 | + | + | EC50(uM) | 50000 | N.A | N.A | 10.1016/j.phrs.2015.07.002 | |
| butanol | 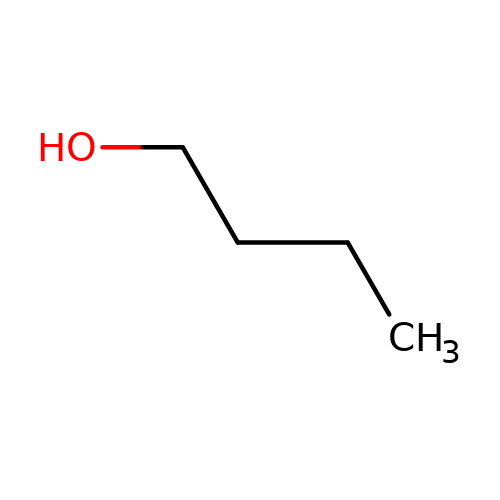 | CCCCO | alcohol | alcohol | 5 | + | + | Kd(uM) | 14700 | Kd(uM) | 14700 | 10.1016/S0028-3908(02)00213-7 | |
| 11S-11-Aminostrychnine | 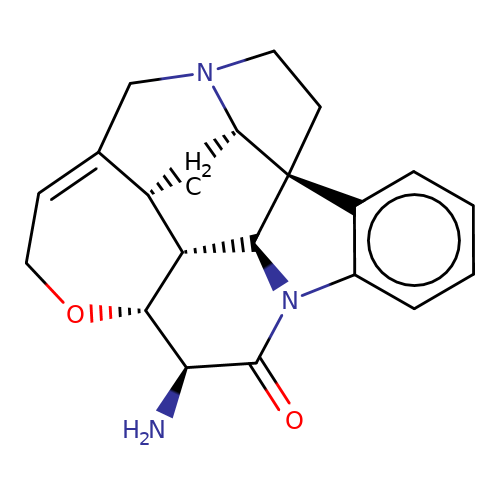 | N[C@@H]1C(=O)N2c3ccccc3[C@@]34CCN5CC6=CCO[C@@H]1[C@@H]([C@H]23)[C@H]6C[C@H]54 | alkaloid | orthosteric | 5 | - | N.A | Ki(uM) | 0.741 | N.A | N.A | 10.1021/acs.jnatprod.9b00180 | |
| N-11S-strychnine-11-ylpropionamide | 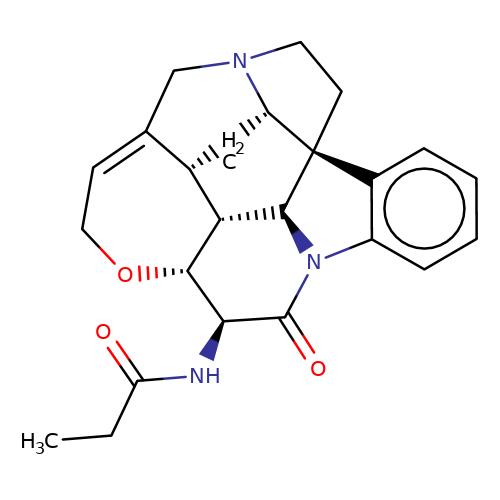 | CCC(=O)N[C@@H]1C(=O)N2c3ccccc3[C@@]34CCN5CC6=CCO[C@@H]1[C@@H]([C@H]23)[C@H]6C[C@H]54 | alkaloid | orthosteric | 5 | - | N.A | Ki(uM) | 0.092 | N.A | N.A | 10.1021/acs.jnatprod.9b00180 | |
| strychnine | 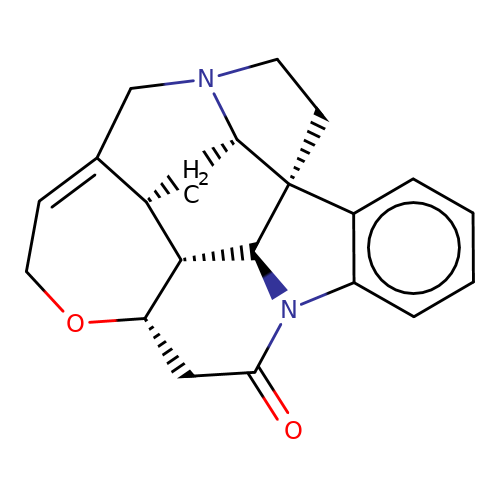 | O=C1C[C@@H]2OCC=C3CN4CC[C@]56c7ccccc7N1[C@H]5[C@H]2[C@H]3C[C@H]46 | alkaloid | orthosteric | 1 | - | - | Ki(uM) | 0.123 | N.A | N.A | 10.1021/acs.jnatprod.9b00180 | |
| gelsemine | 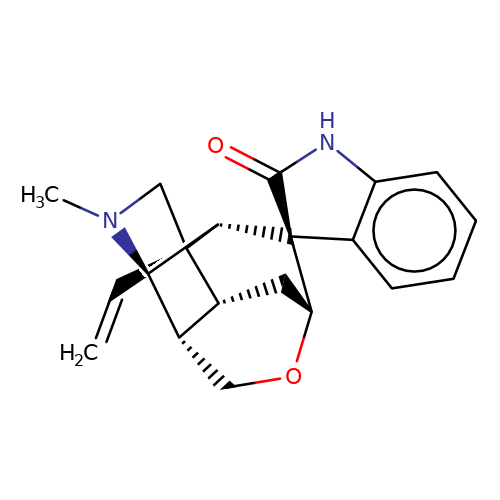 | C=C[C@]12CN(C)[C@@H]3[C@H]4CO[C@H](C[C@H]41)[C@]1(C(=O)Nc4ccccc41)[C@@H]32 | alkaloid | N.A | N.A | + | - | EC50(uM) | 0.59 | IC50(uM) | 36.7 | 10.1111/bph.13507 | |
| glycine | 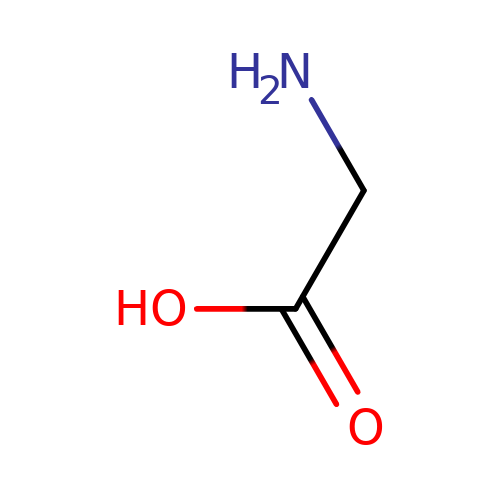 | NCC(=O)O | aminoacid | orthosteric | 1 | + | + | Kd(uM) | 160 | N.A | N.A | 10.1113/jphysiol.2002.037796 | |
| taurine | 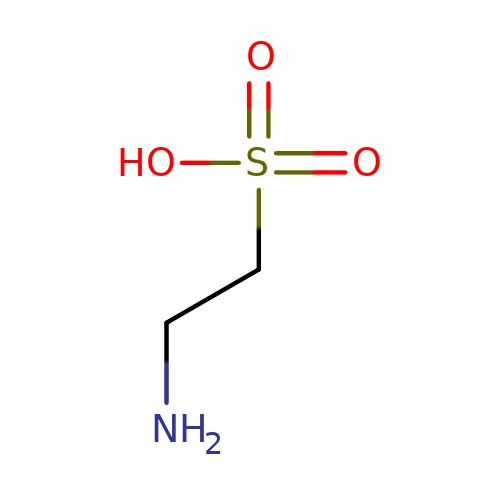 | NCCS(=O)(=O)O | aminoacid | orthosteric | 1 | + | + | Kd(uM) | 460 | N.A | N.A | 10.1113/jphysiol.2002.037796 | |
| beta-alanine | 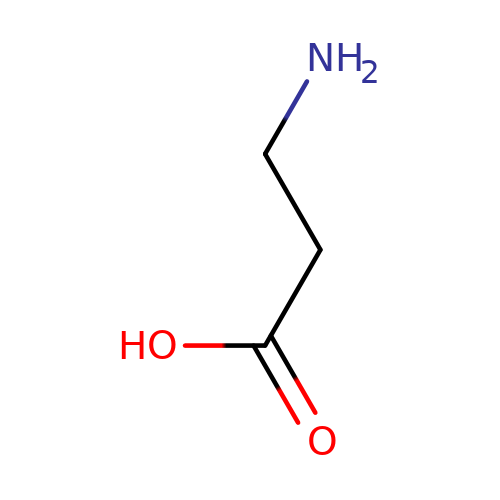 | NCCC(=O)O | aminoacid | orthosteric | 3 | + | + | Kd(uM) | 360 | N.A | N.A | 10.1113/jphysiol.2002.037796 | |
| GABA | 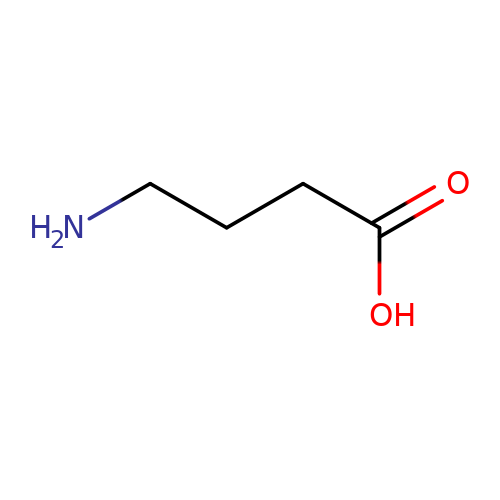 | NCCCC(=O)O | aminoacid | orthosteric | 1 | + | N.A | EC50(uM) | 40000 | N.A | N.A | 10.1111/j.1469-7793.1999.0369t.x | |
| doramectin-b1a | 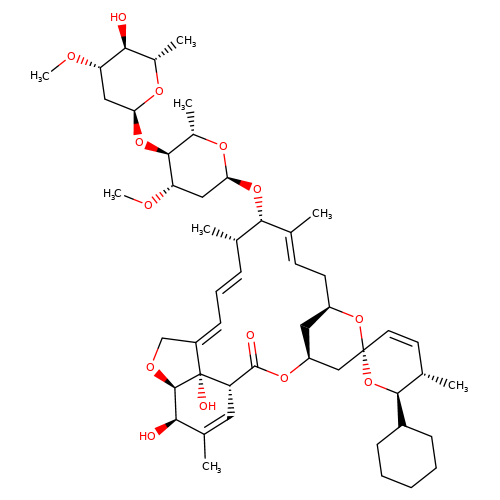 | CO[C@H]1C[C@H](O[C@H]2[C@H](C)O[C@@H](O[C@@H]3/C(C)=C/C[C@@H]4C[C@@H](C[C@]5(C=C[C@H](C)[C@@H](C6CCCCC6)O5)O4)OC(=O)[C@@H]4C=C(C)[C@@H](O)[C@H]5OC/C(=C\C=C\[C@@H]3C)[C@@]45O)C[C@@H]2OC)O[C@@H](C)[C@@H]1O | avermectin | ivermectin | 3 | + | N.A | EC50(uM) | 1.9 | N.A | N.A | 10.1074/jbc.M110.107789 | |
| emamectin-b1a | 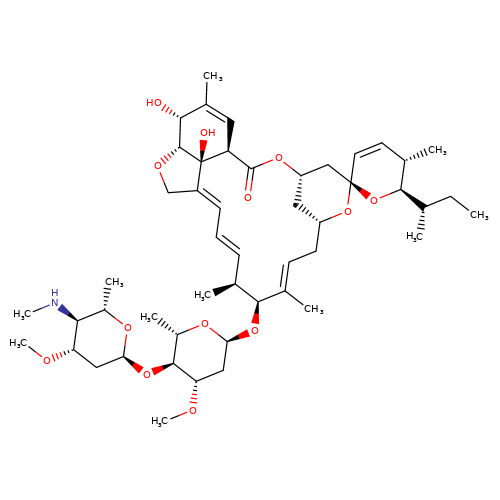 | CC[C@H](C)[C@H]1O[C@]2(C=C[C@@H]1C)C[C@@H]1C[C@@H](C/C=C(\C)[C@@H](O[C@H]3C[C@H](OC)[C@@H](O[C@H]4C[C@H](OC)[C@@H](NC)[C@H](C)O4)[C@H](C)O3)[C@@H](C)/C=C/C=C3\CO[C@@H]4[C@H](O)C(C)=C[C@@H](C(=O)O1)[C@]34O)O2 | avermectin | ivermectin | 3 | + | N.A | EC50(uM) | 1.2 | N.A | N.A | 10.1074/jbc.M110.107789 | |
| eprinomectin | 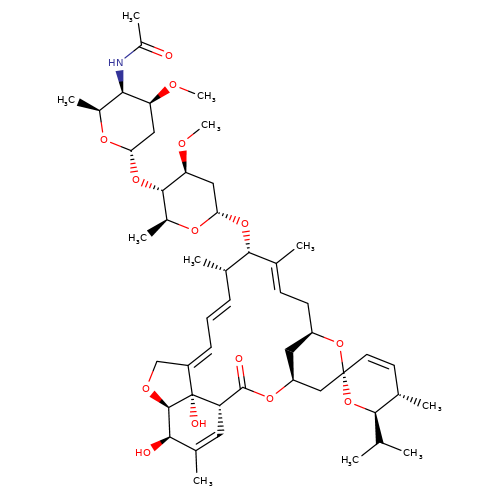 | CO[C@H]1C[C@H](O[C@@H]2/C(C)=C/C[C@@H]3C[C@@H](C[C@]4(C=C[C@H](C)[C@@H](C(C)C)O4)O3)OC(=O)[C@@H]3C=C(C)[C@@H](O)[C@H]4OC/C(=C\C=C\[C@@H]2C)[C@@]34O)O[C@@H](C)[C@@H]1O[C@H]1C[C@H](OC)[C@H](NC(C)=O)[C@H](C)O1 | avermectin | ivermectin | 3 | + | N.A | EC50(uM) | 3.4 | N.A | N.A | 10.1074/jbc.M110.107789 | |
| ivermectin-b1a | 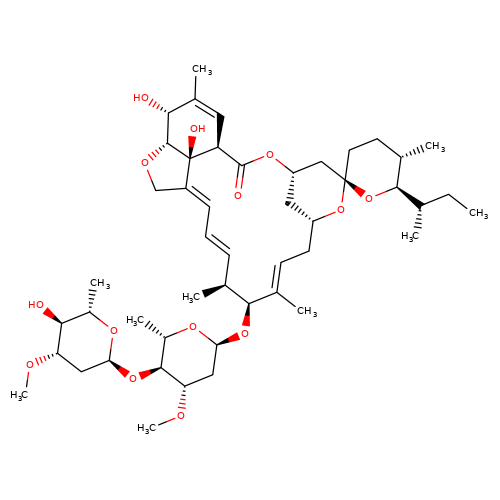 | CC[C@H](C)[C@H]1O[C@]2(CC[C@@H]1C)C[C@@H]1C[C@@H](C/C=C(\C)[C@@H](O[C@H]3C[C@H](OC)[C@@H](O[C@H]4C[C@H](OC)[C@@H](O)[C@H](C)O4)[C@H](C)O3)[C@@H](C)/C=C/C=C3\CO[C@@H]4[C@H](O)C(C)=C[C@@H](C(=O)O1)[C@]34O)O2 | avermectin | ivermectin | 1 | + | N.A | EC50(uM) | 1.7 | N.A | N.A | 10.1074/jbc.M110.107789 | |
| moxidectin | 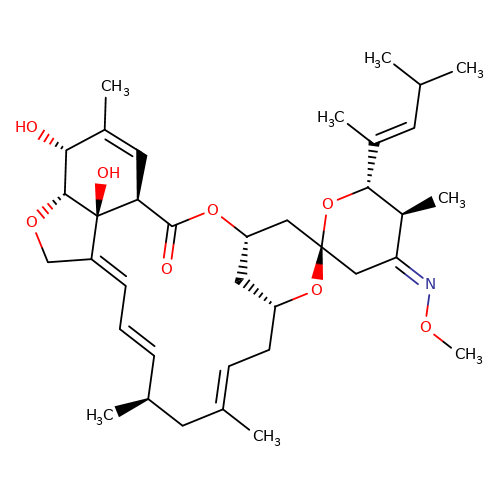 | CO/N=C1\C[C@]2(C[C@@H]3C[C@@H](C/C=C(\C)C[C@@H](C)/C=C/C=C4\CO[C@@H]5[C@H](O)C(C)=C[C@@H](C(=O)O3)[C@]45O)O2)O[C@H](/C(C)=C/C(C)C)[C@H]1C | avermectin | ivermectin | 3 | + | N.A | EC50(uM) | 2.3 | N.A | N.A | 10.1074/jbc.M110.107789 | |
| selamectin | 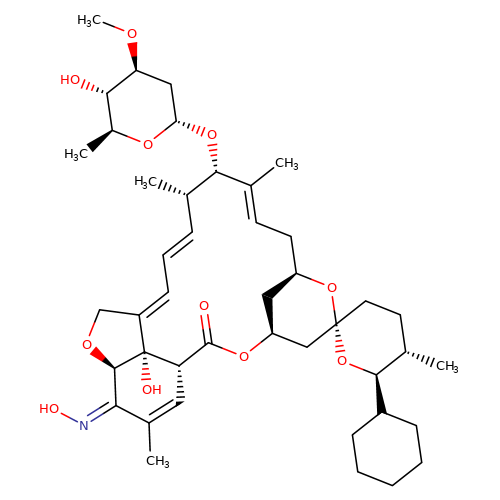 | CO[C@H]1C[C@H](O[C@@H]2/C(C)=C/C[C@@H]3C[C@@H](C[C@]4(CC[C@H](C)[C@@H](C5CCCCC5)O4)O3)OC(=O)[C@@H]3C=C(C)/C(=N/O)[C@H]4OC/C(=C\C=C\[C@@H]2C)[C@@]34O)O[C@@H](C)[C@@H]1O | avermectin | ivermectin | 3 | + | N.A | EC50(uM) | 3.6 | N.A | N.A | 10.1074/JBC.M111.262634 | |
| bilobalide | 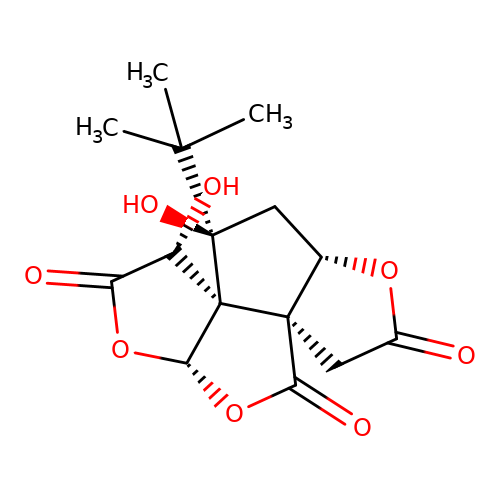 | CC(C)(C)[C@]1(O)C[C@@H]2OC(=O)C[C@@]23C(=O)O[C@@H]2OC(=O)[C@H](O)[C@@]231 | bilabolide | pore | 3 | - | N.A | IC50(uM) | 19.6 | N.A | N.A | 10.1111/j.1471-4159.2006.03875.x | |
| NV-31 | 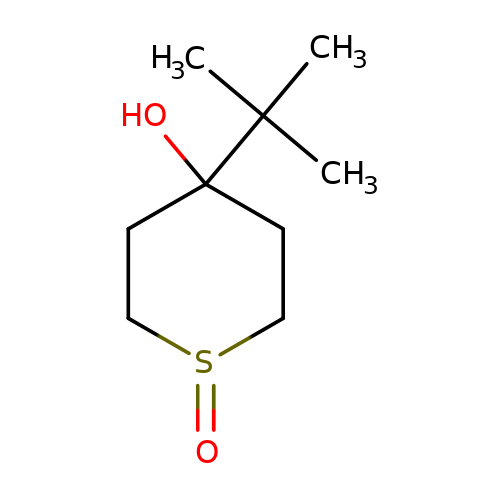 | CC(C)(C)C1(O)CCS(=O)CC1 | bilabolide | ivermectin | 3 | + | 0 | EC50(uM) | 0.17 | Maximum_effect(%) | 100 | 10.1016/j.neulet.2008.02.022 | |
| THC | 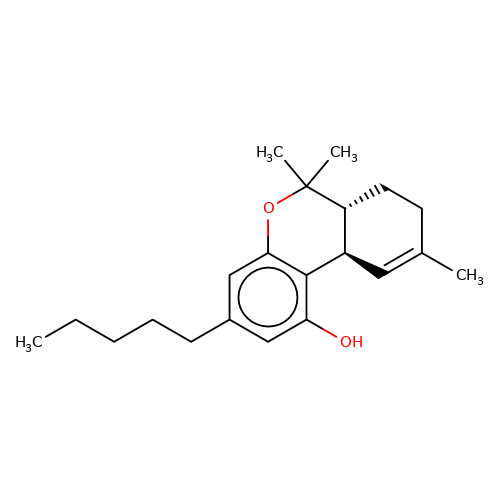 | CCCCCc1cc(O)c2c(c1)OC(C)(C)[C@@H]1CCC(C)=C[C@@H]21 | cannabinoid | N.A | N.A | + | N.A | EC50(uM) | 0.086 | N.A | N.A | 10.1124/mol.105.019174 | |
| anandamide | 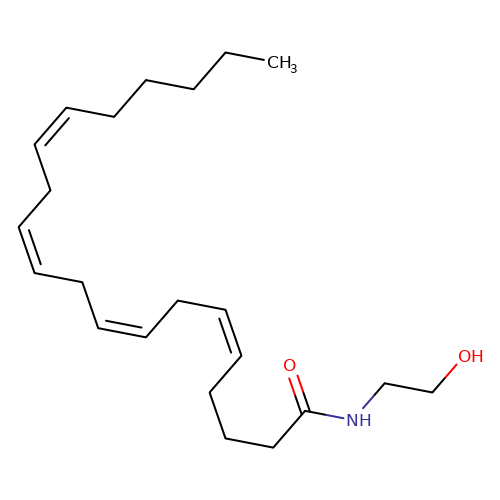 | CCCCC/C=C\C/C=C\C/C=C\C/C=C\CCCC(=O)NCCO | cannabinoid | N.A | N.A | + | 0 | EC50(uM) | 0.038 | IC50(uM) | Inactive | 10.1016/j.bcp.2008.07.037 | |
| HU-210 | 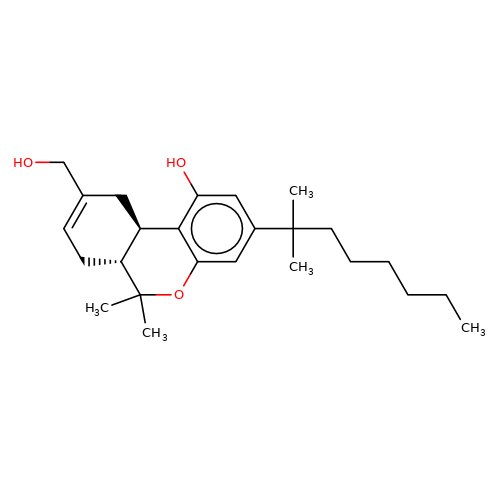 | CCCCCCC(C)(C)c1cc(O)c2c(c1)OC(C)(C)[C@@H]1CC=C(CO)C[C@@H]21 | cannabinoid | N.A | N.A | + | - | EC50(uM) | 0.27 | IC50(uM) | 0.05 | 10.1016/j.bcp.2008.07.037 | |
| HU-308 | 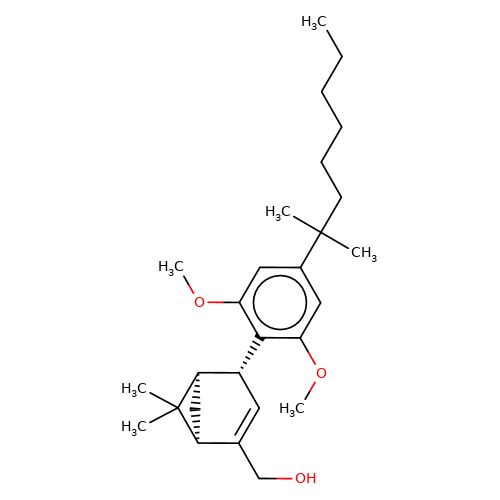 | CCCCCCC(C)(C)c1cc(OC)c([C@@H]2C=C(CO)[C@@H]3C[C@H]2C3(C)C)c(OC)c1 | cannabinoid | N.A | N.A | 0 | - | EC50(uM) | Inactive | IC50(uM) | 0.097 | 10.1016/j.bcp.2008.07.037 | |
| N-Arachidonoyl-glycine | 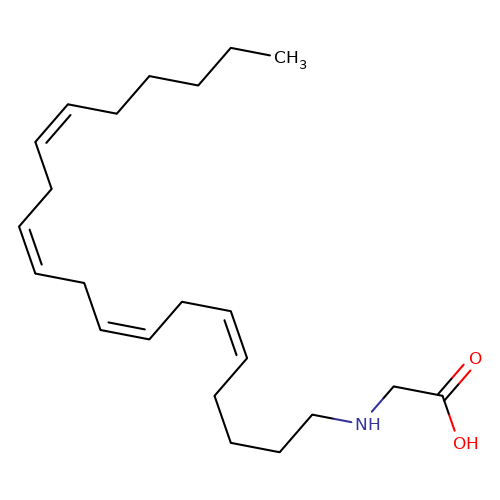 | CCCCC/C=C\C/C=C\C/C=C\C/C=C\CCCCNCC(=O)O | cannabinoid | N.A | N.A | 0 | - | EC50(uM) | Inactive | IC50(uM) | 1.32 | 10.1016/j.bcp.2008.07.037 | |
| WIN-55212-3 | 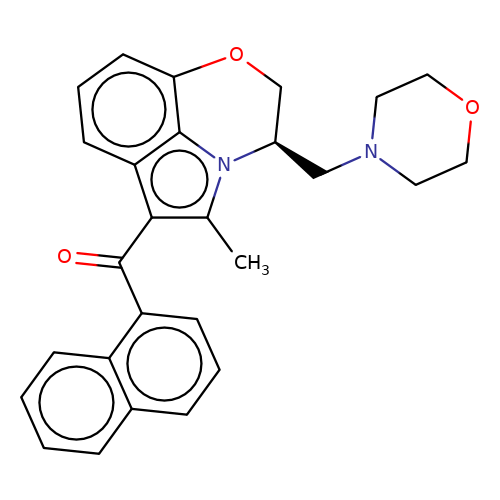 | Cc1c(C(=O)c2cccc3ccccc23)c2cccc3c2n1[C@H](CN1CCOCC1)CO3 | cannabinoid | N.A | N.A | 0 | - | EC50(uM) | Inactive | IC50(uM) | 0.086 | 10.1016/j.bcp.2008.07.037 | |
| 2-Arachidonoylglycerol | 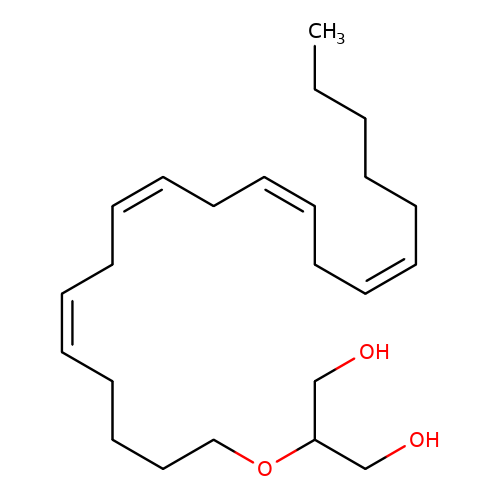 | CCCCC/C=C\C/C=C\C/C=C\C/C=C\CCCCOC(CO)CO | cannabinoid | N.A | N.A | - | - | Maximum_effect(%)nosubtype | 40 | Maximum_effect(%)nosubtype | 40 | 10.1523/JNEUROSCI.0977-05.2005 | |
| DH-CBD | 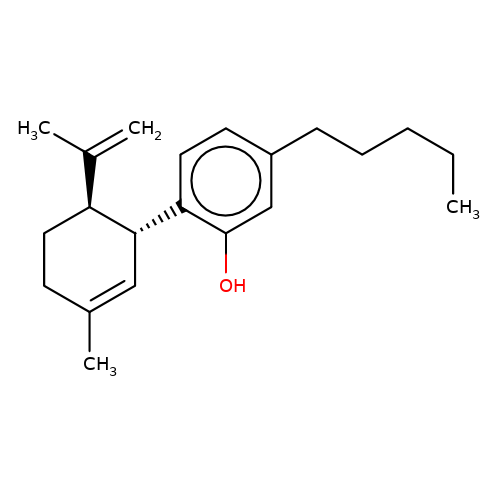 | C=C(C)[C@@H]1CCC(C)=C[C@H]1c1ccc(CCCCC)cc1O | cannabinoid | N.A | N.A | N.A | + | N.A | N.A | Mean_potentiation(%) | 989 | 10.1084/jem.20120242 | |
| CBD | 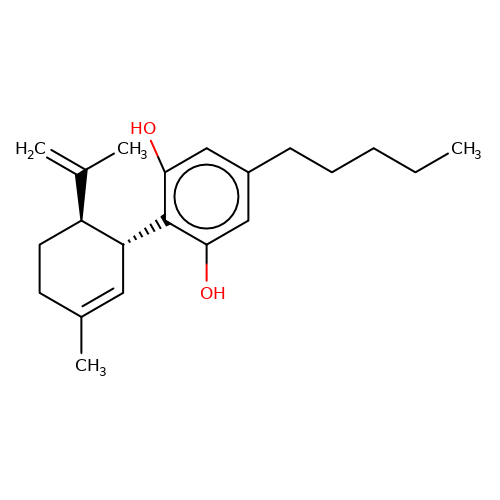 | C=C(C)[C@@H]1CCC(C)=C[C@H]1c1c(O)cc(CCCCC)cc1O | cannabinoid | N.A | N.A | N.A | + | N.A | N.A | Mean_potentiation(%) | 491 | 10.1084/jem.20120242 | |
| DD-CBD | 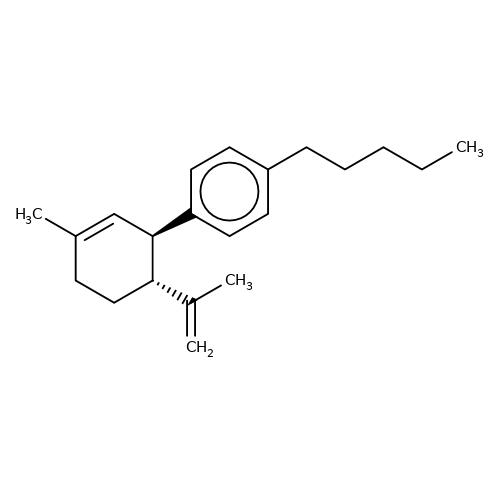 | C=C(C)[C@@H]1CCC(C)=C[C@H]1c1ccc(CCCCC)cc1 | cannabinoid | N.A | N.A | N.A | 0 | N.A | N.A | Mean_potentiation(%) | Inactive | 10.1084/jem.20120242 | |
| N-Arachidonoyl-serine | 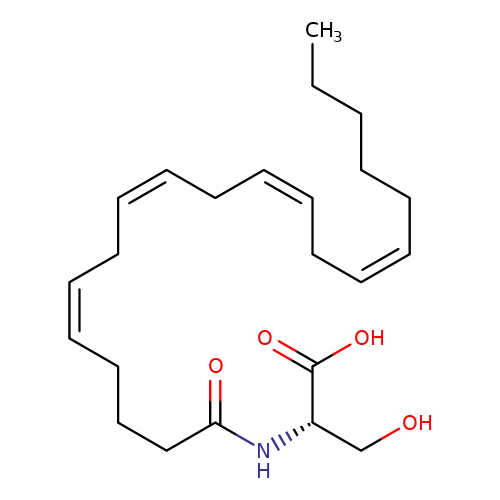 | CCCCC/C=C\C/C=C\C/C=C\C/C=C\CCCC(=O)N[C@@H](CO)C(=O)O | cannabinoid | N.A | N.A | + | - | Mean_potentiation(%) | 175 | Mean_potentiation(%) | 50 | 10.1371/journal.pone.0023886 | |
| Ajulemic-acid | 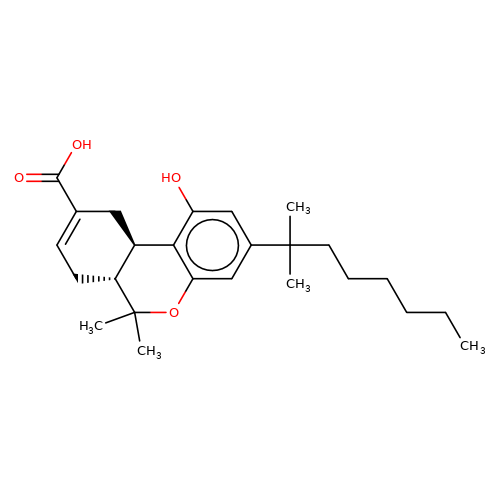 | CCCCCCC(C)(C)c1cc(O)c2c(c1)OC(C)(C)[C@@H]1CC=C(C(=O)O)C[C@@H]21 | cannabinoid | N.A | N.A | + | N.A | EC50(uM) | 9.7 | N.A | N.A | 10.1007/s00210-008-0366-8 | |
| cyanotriphenylborate | 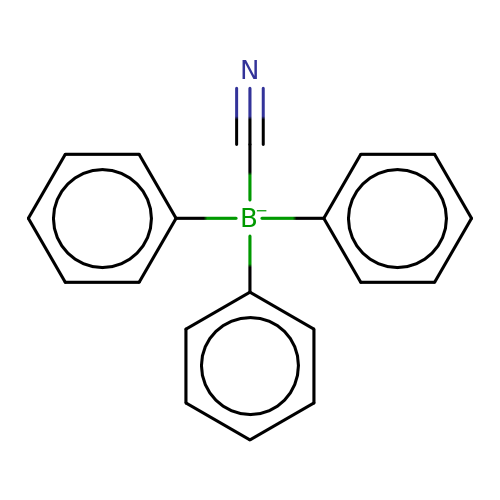 | N#C[B-](c1ccccc1)(c1ccccc1)c1ccccc1 | cyanotriphenylborate | pore | 3 | - | N.A | IC50(uM) | 1.3 | N.A | N.A | 10.1073/pnas.91.19.8950 | |
| nicardipine | 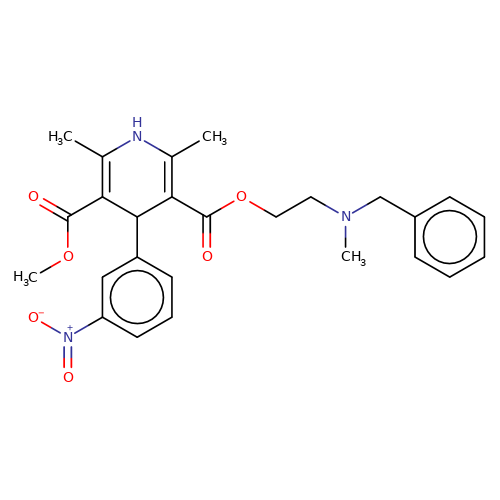 | COC(=O)C1=C(C)NC(C)=C(C(=O)OCCN(C)Cc2ccccc2)C1c1cccc([N+](=O)[O-])c1 | dihydropyridine | pore | 3 | - | N.A | IC50(uM) | 3.3 | IC50(uM) | 29.2 | 10.1016/J.NEUROPHARM.2008.07.001 | |
| nifedipine | 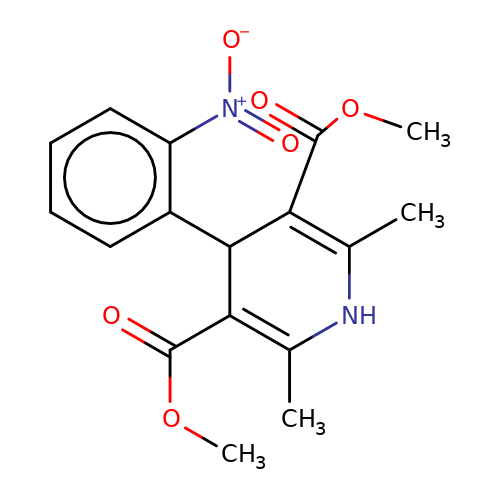 | COC(=O)C1=C(C)NC(C)=C(C(=O)OC)C1c1ccccc1[N+](=O)[O-] | dihydropyridine | pore | 3 | - | N.A | IC50(uM) | 10.6 | IC50(uM) | 30 | 10.1016/J.NEUROPHARM.2008.07.001 | |
| nitrenpidine | 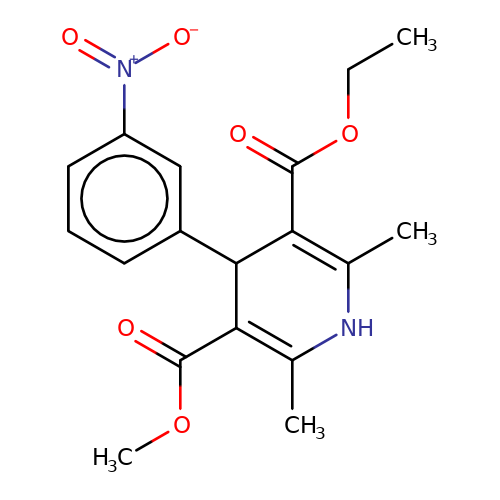 | CCOC(=O)C1=C(C)NC(C)=C(C(=O)OC)C1c1cccc([N+](=O)[O-])c1 | dihydropyridine | pore | 3 | - | N.A | Mean_potentiation(%) | 17 | N.A | N.A | 10.1046/j.0953-816x.2001.01599.x | |
| enflurane | 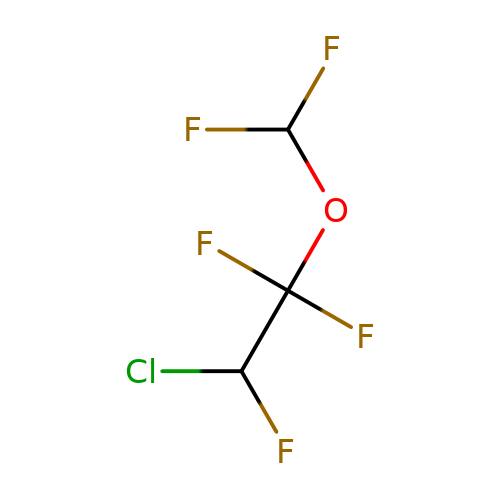 | FC(F)OC(F)(F)C(F)Cl | general_anesthetic | N.A | N.A | + | N.A | EC50(uM) | 500 | N.A | N.A | 10.1038/sj.bjp.0703087 | |
| halothane | 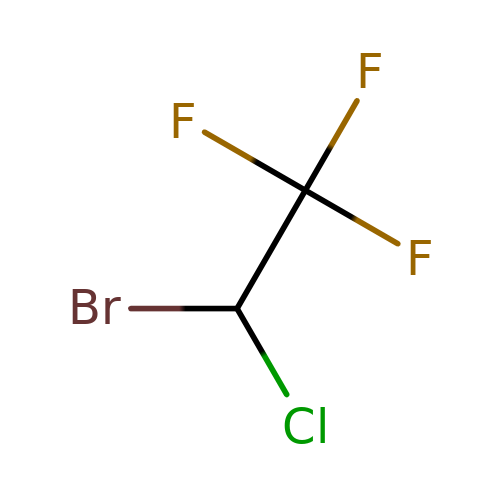 | FC(F)(F)C(Cl)Br | general_anesthetic | N.A | N.A | + | N.A | EC50(uM) | 180 | N.A | N.A | 10.1111/j.1476-5381.1996.tb15430.x | |
| isoflurane | 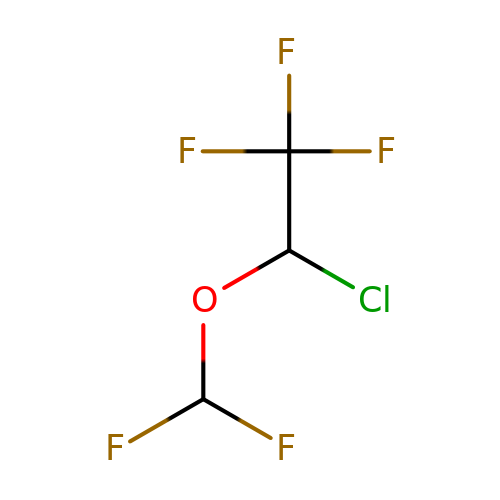 | FC(F)OC(Cl)C(F)(F)F | general_anesthetic | N.A | N.A | + | N.A | EC50(uM) | 270 | N.A | N.A | 10.1038/sj.bjp.0703087 | |
| sevoflurane | 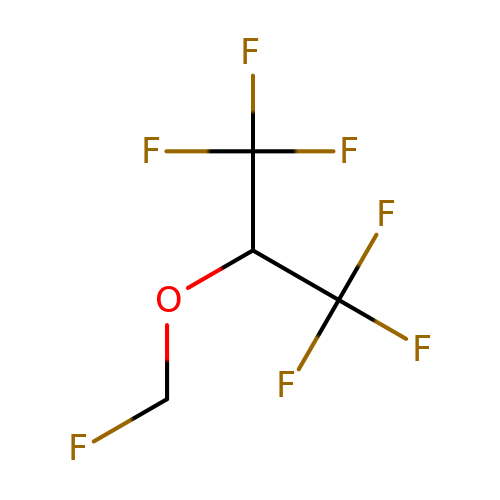 | FCOC(C(F)(F)F)C(F)(F)F | general_anesthetic | N.A | N.A | + | N.A | EC50(uM) | 360 | N.A | N.A | 10.1038/sj.bjp.0703087 | |
| methoxyflurane | 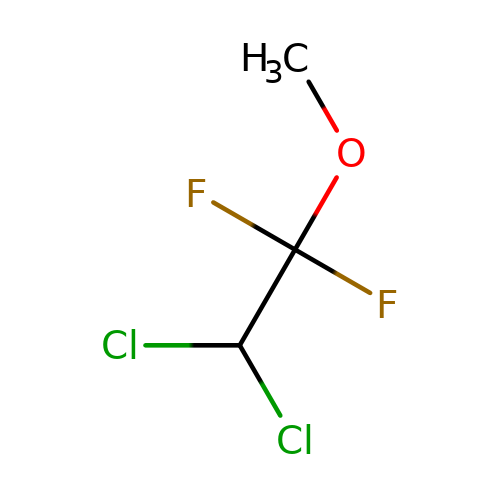 | COC(F)(F)C(Cl)Cl | general_anesthetic | N.A | N.A | + | N.A | EC50(uM) | 540 | N.A | N.A | 10.1038/sj.bjp.0703087 | |
| ginkgolic-acid | 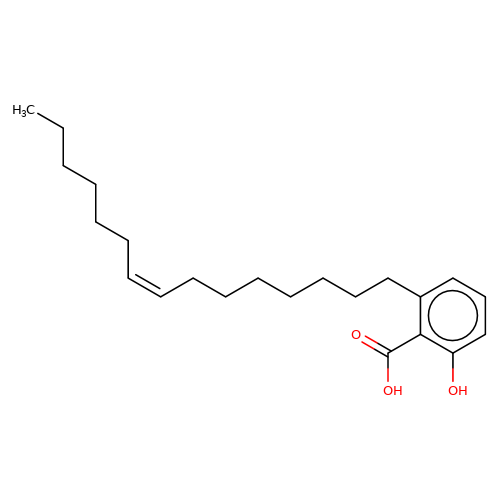 | CCCCCC/C=C\CCCCCCCc1cccc(O)c1C(=O)O | ginkgolic-acid | N.A | N.A | + | + | Maximum_effect(%) | 182 | Maximum_effect(%) | 100 | 10.3389/fnmol.2015.00064 | |
| ginkgolide-A | 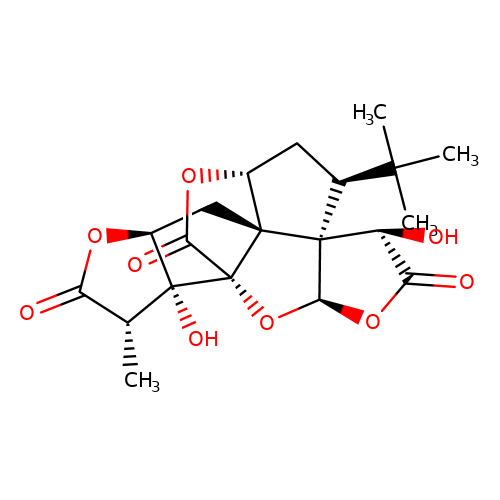 | C[C@@H]1C(=O)O[C@H]2C[C@@]34[C@H]5C[C@@H](C(C)(C)C)[C@]36[C@@H](OC(=O)[C@@H]6O)O[C@@]4(C(=O)O5)[C@]21O | ginkgolide | pore | 3 | - | N.A | IC50(uM) | 1.9 | N.A | N.A | 10.1021/jm070003n | |
| ginkgolide-B | 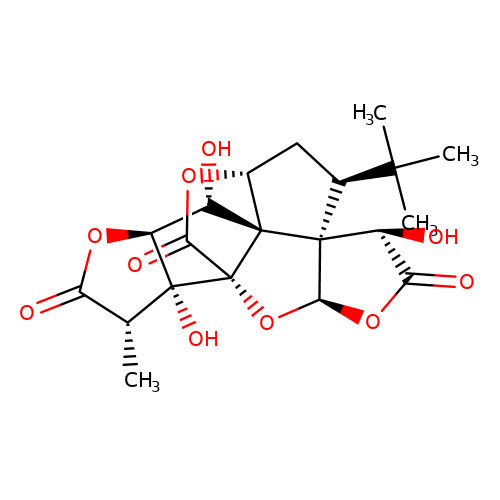 | C[C@@H]1C(=O)O[C@H]2[C@H](O)[C@@]34[C@H]5C[C@@H](C(C)(C)C)[C@]36[C@@H](OC(=O)[C@@H]6O)O[C@@]4(C(=O)O5)[C@@]12O | ginkgolide | pore | 3 | - | N.A | IC50(uM) | 2.9 | N.A | N.A | 10.1021/jm070003n | |
| ginkgolide-C | 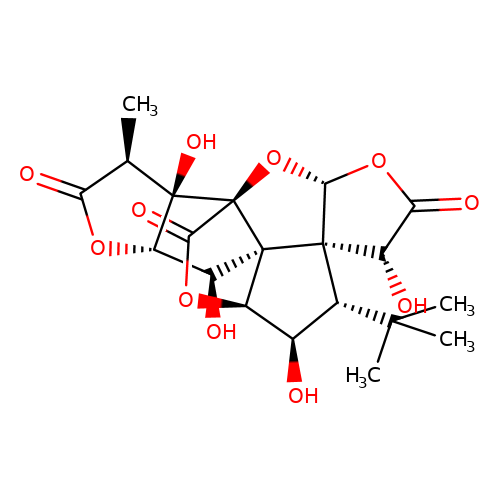 | C[C@@H]1C(=O)O[C@H]2[C@H](O)[C@]34[C@@H]5OC(=O)[C@]3(O[C@@H]3OC(=O)[C@H](O)[C@@]34[C@H](C(C)(C)C)[C@H]5O)[C@@]12O | ginkgolide | pore | 3 | - | N.A | IC50(uM) | 4.7 | N.A | N.A | 10.1021/jm070003n | |
| ginkgolide-J | 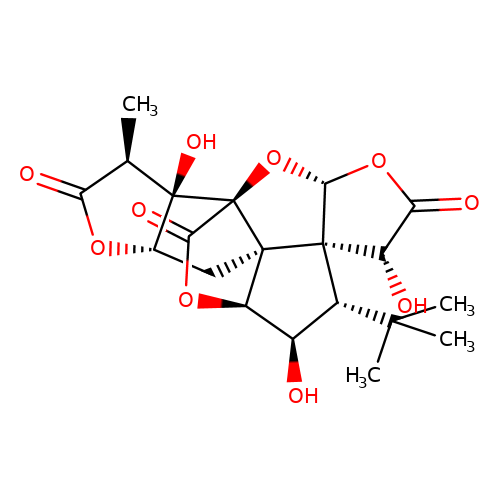 | C[C@@H]1C(=O)O[C@H]2C[C@]34[C@@H]5OC(=O)[C@]3(O[C@@H]3OC(=O)[C@H](O)[C@@]34[C@H](C(C)(C)C)[C@H]5O)[C@]21O | ginkgolide | pore | 3 | - | N.A | IC50(uM) | 7.1 | N.A | N.A | 10.1021/jm070003n | |
| ginkgolide-M | 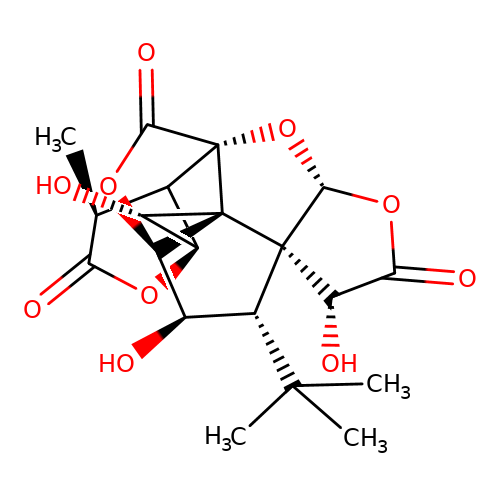 | C[C@@H]1C(=O)O[C@@H]2C1[C@@]13O[C@@H]4OC(=O)[C@H](O)[C@@]45[C@H](C(C)(C)C)[C@@H](O)[C@@H](OC1=O)[C@@]35[C@H]2O | ginkgolide | pore | 3 | - | N.A | IC50(uM) | 0.78 | N.A | N.A | 10.1021/jm070003n | |
| ginkgolide-X | 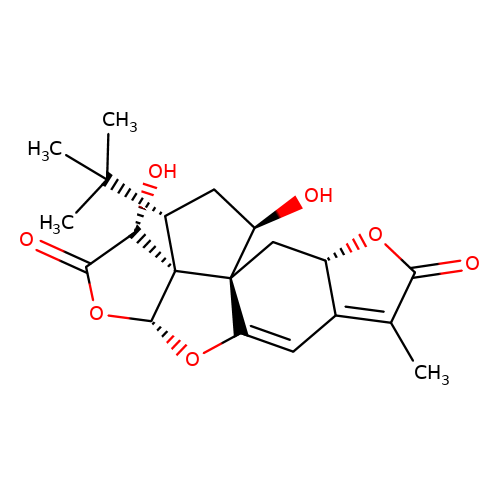 | CC1=C2C=C3O[C@@H]4OC(=O)[C@H](O)[C@@]45[C@H](C(C)(C)C)C[C@@H](O)[C@@]35C[C@@H]2OC1=O | ginkgolide | pore | 3 | - | N.A | IC50(uM) | 1.4 | N.A | N.A | 10.1074/jbc.M109.079319 | |
| Jensen-6 | 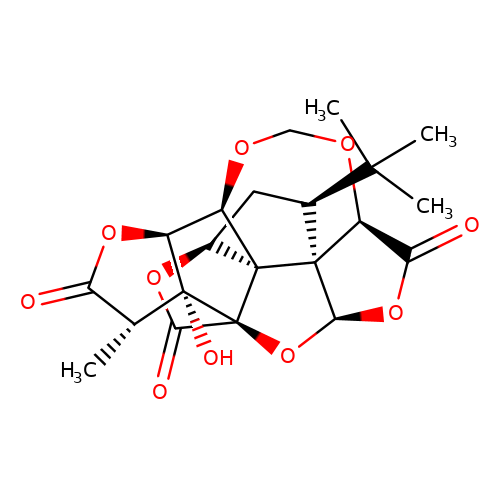 | C[C@@H]1C(=O)O[C@H]2[C@H]3OCO[C@H]4C(=O)O[C@H]5O[C@@]6(C(=O)O[C@H]7C[C@@H](C(C)(C)C)[C@@]54[C@]736)[C@@]12O | ginkgolide | pore | 5 | - | N.A | IC50(uM) | 12 | N.A | N.A | 10.1021/jm070003n | |
| Jensen-7 | 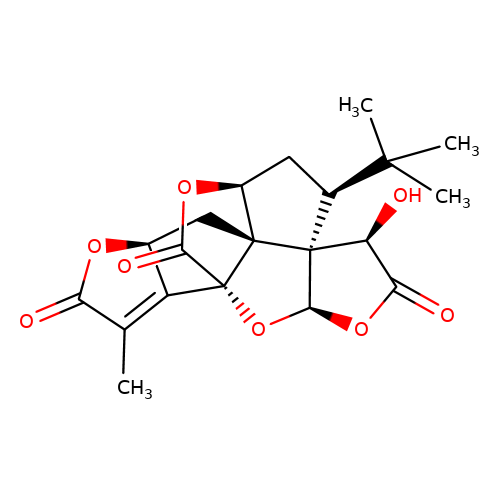 | CC1=C2[C@H](C[C@@]34[C@@H]5C[C@@H](C(C)(C)C)[C@]36[C@@H](OC(=O)[C@@H]6O)O[C@@]24C(=O)O5)OC1=O | ginkgolide | pore | 5 | - | N.A | IC50(uM) | 6.2 | N.A | N.A | 10.1021/jm070003n | |
| Jensen-12 | 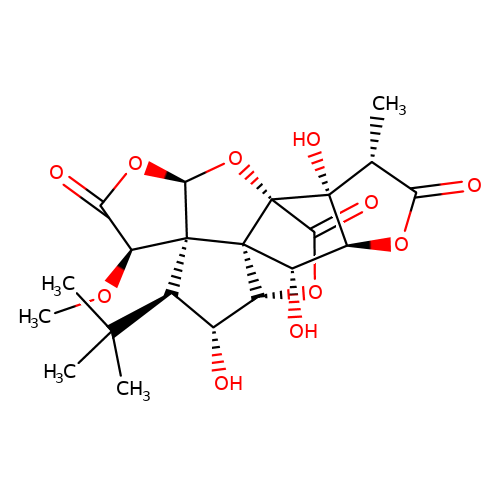 | CO[C@H]1C(=O)O[C@H]2O[C@]34C(=O)O[C@@H]5[C@H](O)[C@@H](C(C)(C)C)[C@@]21[C@@]53[C@@H](O)[C@@H]1OC(=O)[C@@H](C)[C@@]14O | ginkgolide | pore | 5 | - | N.A | IC50(uM) | 9.8 | N.A | N.A | 10.1021/jm070003n | |
| Jensen-27 | 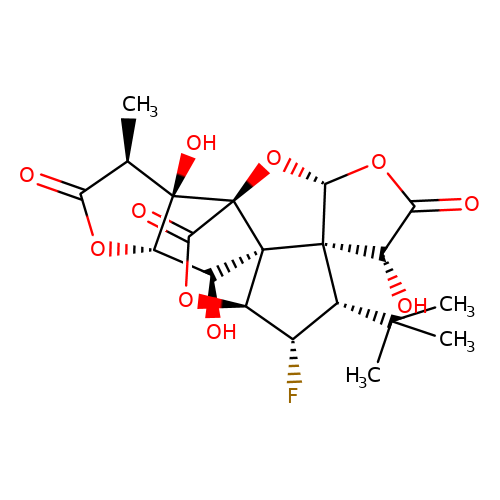 | C[C@@H]1C(=O)O[C@H]2[C@H](O)[C@]34[C@@H]5OC(=O)[C@]3(O[C@@H]3OC(=O)[C@H](O)[C@@]34[C@H](C(C)(C)C)[C@@H]5F)[C@@]12O | ginkgolide | pore | 5 | - | N.A | IC50(uM) | 7.3 | N.A | N.A | 10.1021/jm070003n | |
| Jensen-28 | 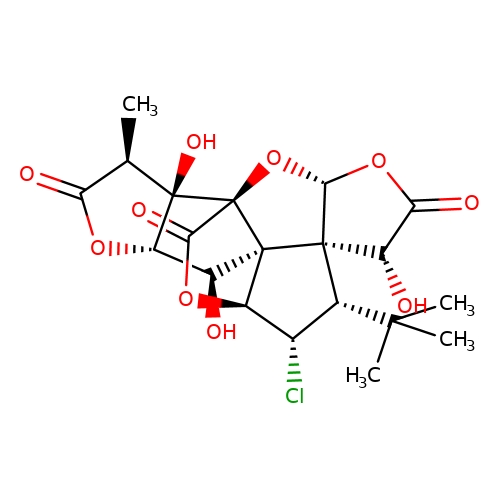 | C[C@@H]1C(=O)O[C@H]2[C@H](O)[C@]34[C@@H]5OC(=O)[C@]3(O[C@@H]3OC(=O)[C@H](O)[C@@]34[C@H](C(C)(C)C)[C@@H]5Cl)[C@@]12O | ginkgolide | pore | 5 | - | N.A | IC50(uM) | 7.6 | N.A | N.A | 10.1021/jm070003n | |
| Jensen-29 | 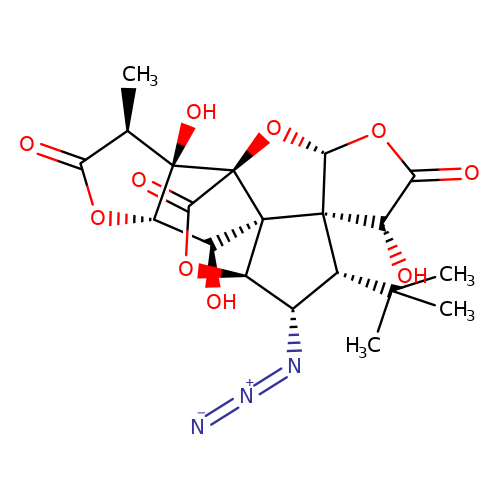 | C[C@@H]1C(=O)O[C@H]2[C@H](O)[C@]34[C@@H]5OC(=O)[C@]3(O[C@@H]3OC(=O)[C@H](O)[C@@]34[C@H](C(C)(C)C)[C@@H]5N=[N+]=[N-])[C@@]12O | ginkgolide | pore | 5 | - | N.A | IC50(uM) | 17 | N.A | N.A | 10.1021/jm070003n | |
| alpha-cortol | 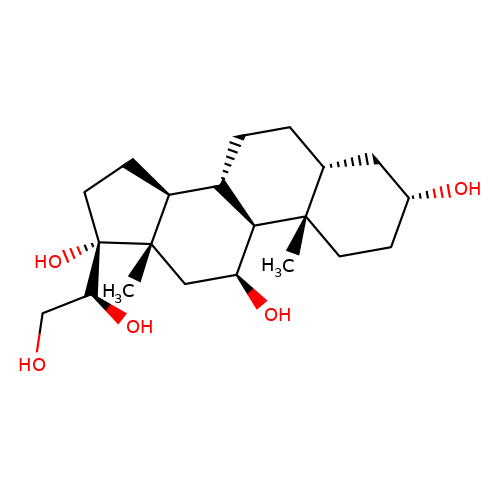 | C[C@]12CC[C@@H](O)C[C@H]1CC[C@@H]1[C@@H]2[C@@H](O)C[C@@]2(C)[C@H]1CC[C@]2(O)[C@@H](O)CO | glucocortico_steroid | N.A | N.A | 0 | N.A | EC50(uM) | Inactive | N.A | N.A | 10.1093/bja/aeh125 | |
| 20alpha-dihydrocortisol | 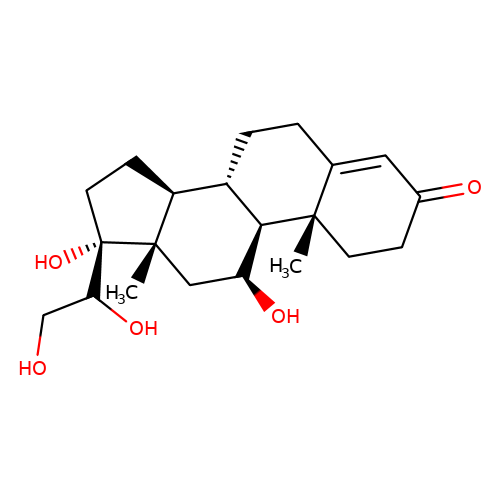 | C[C@]12C[C@H](O)[C@H]3[C@@H](CCC4=CC(=O)CC[C@@]43C)[C@@H]1CC[C@]2(O)C(O)CO | glucocortico_steroid | N.A | N.A | 0 | N.A | EC50(uM) | Inactive | N.A | N.A | 10.1093/bja/aeh125 | |
| hydrocortisone | 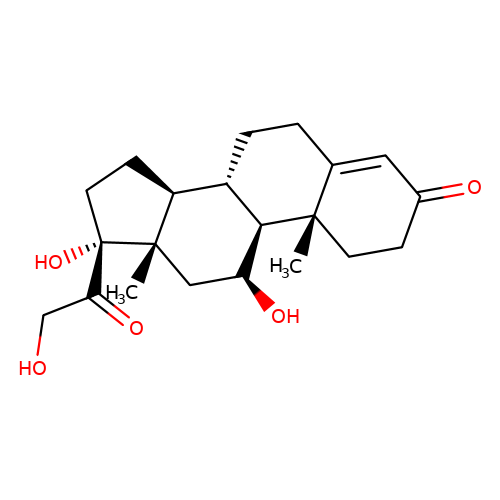 | C[C@]12C[C@H](O)[C@H]3[C@@H](CCC4=CC(=O)CC[C@@]43C)[C@@H]1CC[C@]2(O)C(=O)CO | glucocortico_steroid | N.A | N.A | 0 | N.A | EC50(uM) | Inactive | N.A | N.A | 10.1093/bja/aeh125 | |
| AP5 | 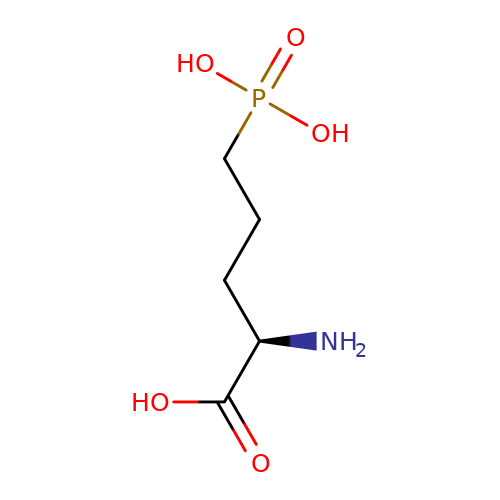 | N[C@H](CCCP(=O)(O)O)C(=O)O | glutamate | N.A | N.A | + | N.A | EC50(uM) | 49.4 | N.A | N.A | 10.1038/nn.2633 | |
| L-glutamate | 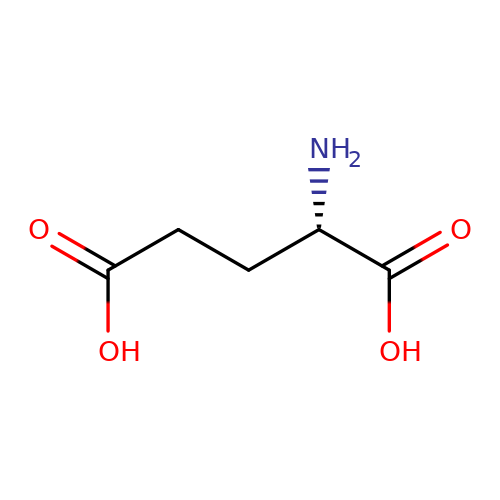 | N[C@@H](CCC(=O)O)C(=O)O | glutamate | N.A | N.A | + | N.A | EC50(uM) | 19.9 | N.A | N.A | 10.1038/nn.2633 | |
| CNQX | 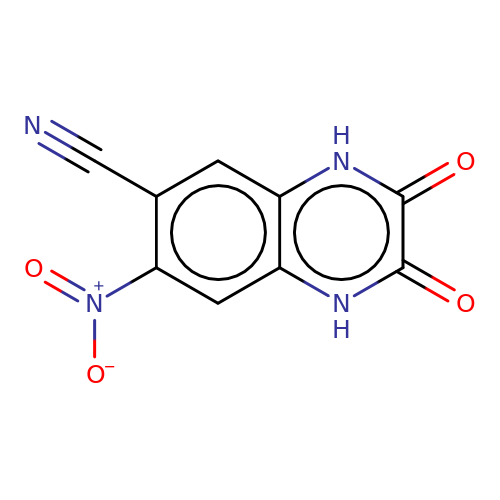 | N#Cc1cc2[nH]c(=O)c(=O)[nH]c2cc1[N+](=O)[O-] | glutamate | N.A | N.A | - | N.A | Maximum_effect(%) | 73 | N.A | N.A | 10.1038/nn.2633 | |
| NBQX | 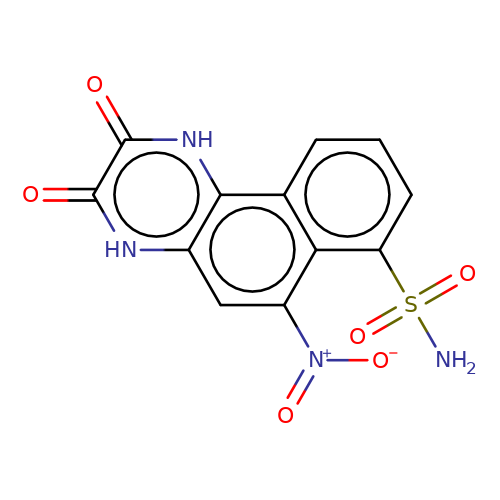 | NS(=O)(=O)c1cccc2c1c([N+](=O)[O-])cc1[nH]c(=O)c(=O)[nH]c12 | glutamate | N.A | N.A | - | N.A | Maximum_effect(%) | 52 | N.A | N.A | 10.1038/nn.2633 | |
| L-aspartate | 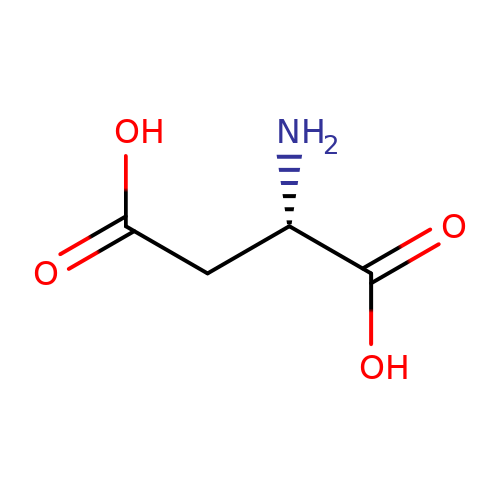 | N[C@@H](CC(=O)O)C(=O)O | glutamate | N.A | N.A | + | N.A | Maximum_effect(%) | 167 | N.A | N.A | 10.1038/nn.2633 | |
| kainate | 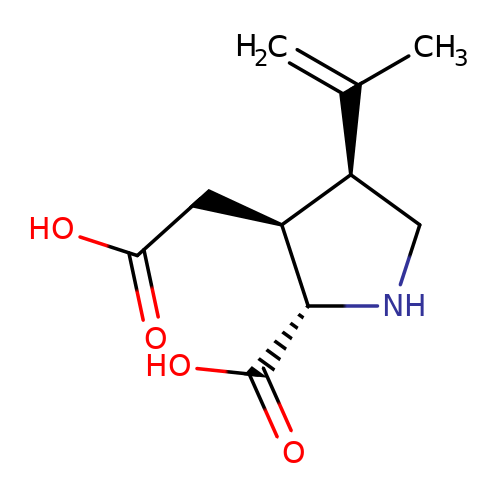 | C=C(C)[C@H]1CN[C@H](C(=O)O)[C@H]1CC(=O)O | glutamate | N.A | N.A | + | N.A | Maximum_effect(%) | 218 | N.A | N.A | 10.1038/nn.2633 | |
| kynurenic-acid | 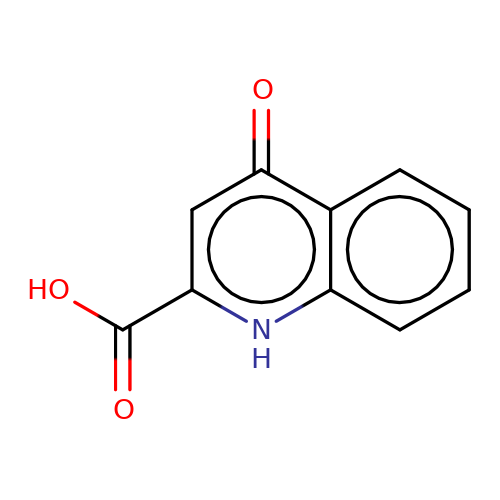 | O=C(O)c1cc(=O)c2ccccc2[nH]1 | glutamate | N.A | N.A | + | N.A | Maximum_effect(%) | 164 | N.A | N.A | 10.1038/nn.2633 | |
| NMDA | 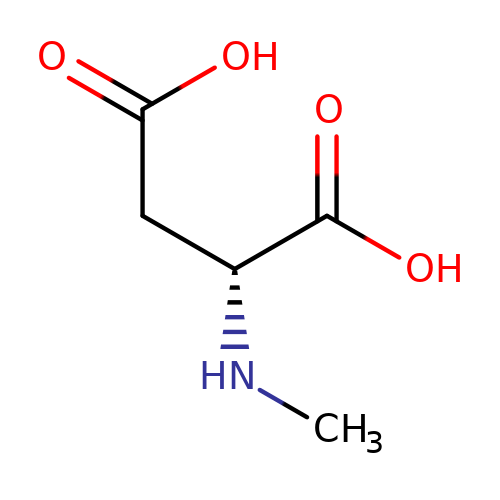 | CN[C@H](CC(=O)O)C(=O)O | glutamate | N.A | N.A | + | N.A | Maximum_effect(%) | 189 | N.A | N.A | 10.1038/nn.2633 | |
| quisqualate | 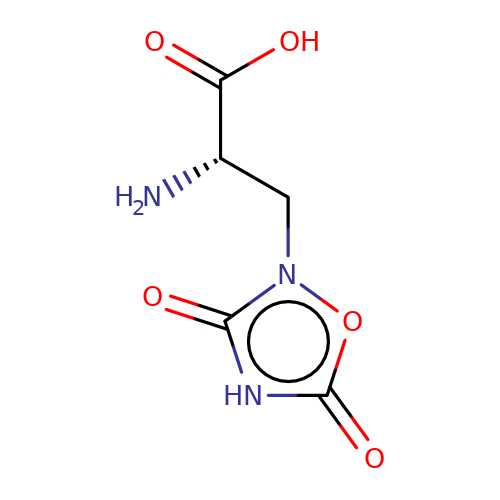 | N[C@@H](Cn1oc(=O)[nH]c1=O)C(=O)O | glutamate | N.A | N.A | + | N.A | Maximum_effect(%) | 203 | N.A | N.A | 10.1038/nn.2633 | |
| alphaxalone | 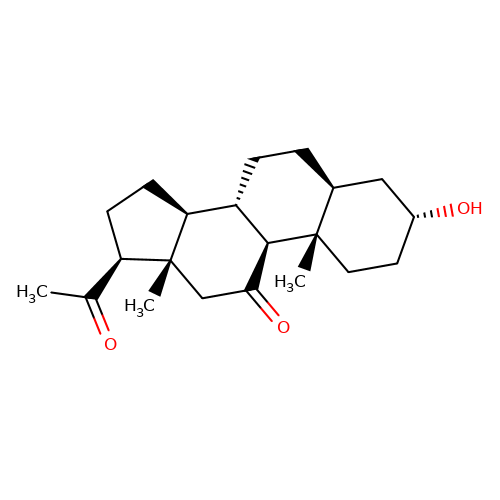 | CC(=O)[C@H]1CC[C@H]2[C@@H]3CC[C@H]4C[C@H](O)CC[C@]4(C)[C@H]3C(=O)C[C@]12C | neurosteroid | (+)-neurosteroid | 5 | + | N.A | EC50(uM) | 28 | N.A | N.A | 10.1093/bja/aeh125 | |
| minaxolone | 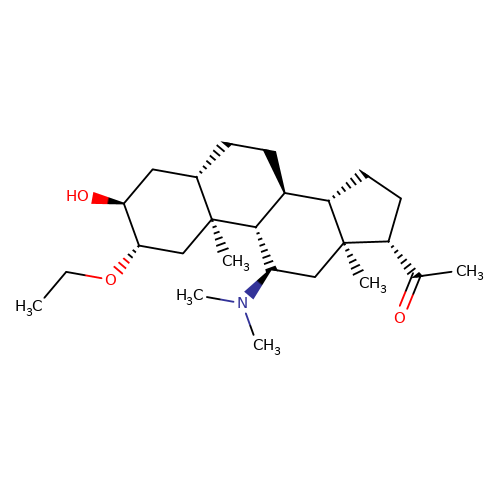 | CCO[C@H]1C[C@@]2(C)[C@@H](CC[C@H]3[C@@H]4CC[C@H](C(C)=O)[C@@]4(C)C[C@@H](N(C)C)[C@@H]32)C[C@@H]1O | neurosteroid | (+)-neurosteroid | 5 | + | N.A | EC50(uM) | 13 | N.A | N.A | 10.1093/bja/aeh125 | |
| ORG-20599 | 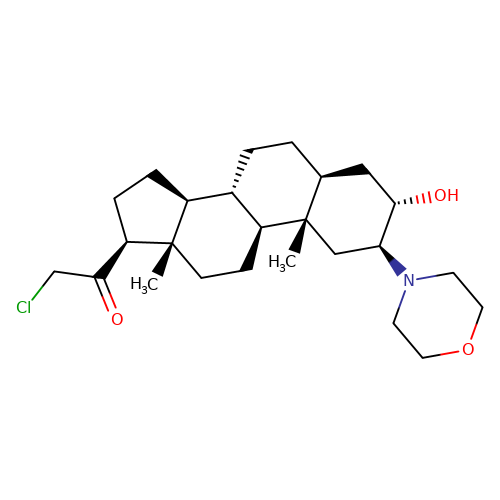 | C[C@]12C[C@H](N3CCOCC3)[C@@H](O)C[C@@H]1CC[C@@H]1[C@@H]2CC[C@]2(C)[C@@H](C(=O)CCl)CC[C@@H]12 | neurosteroid | (+)-neurosteroid | 5 | + | N.A | EC50(uM) | 23 | N.A | N.A | 10.1093/bja/aeh125 | |
| Pregnenolone-acetate | 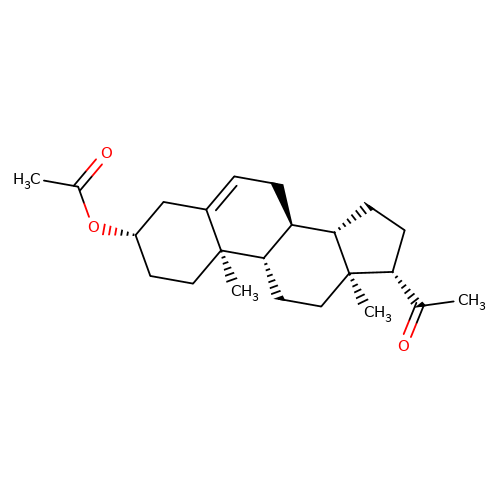 | CC(=O)O[C@H]1CC[C@@]2(C)C(=CC[C@H]3[C@@H]4CC[C@H](C(C)=O)[C@@]4(C)CC[C@@H]32)C1 | neurosteroid | (+)-neurosteroid | 5 | + | N.A | EC50(uM) | 1.1 | N.A | N.A | 10.1016/S0028-3908(01)00071-5 | |
| pregnenolone | 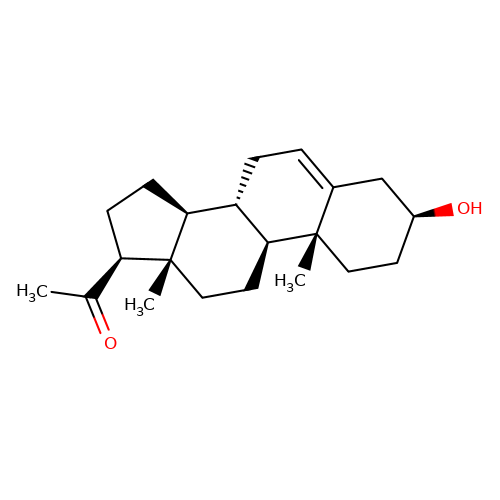 | CC(=O)[C@H]1CC[C@H]2[C@@H]3CC=C4C[C@@H](O)CC[C@]4(C)[C@H]3CC[C@]12C | neurosteroid | (+)-neurosteroid | 5 | + | N.A | EC50(uM) | 1.4 | N.A | N.A | 10.1016/S0028-3908(01)00071-5 | |
| progesterone | 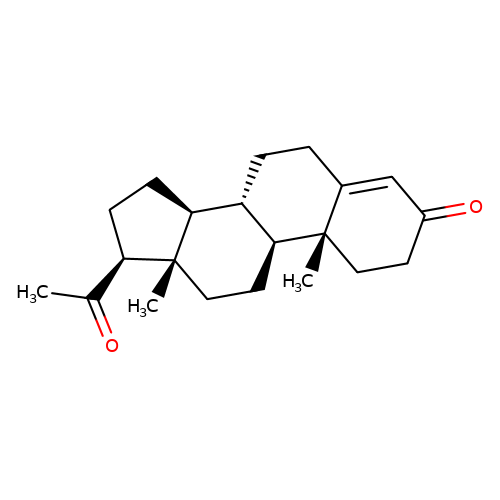 | CC(=O)[C@H]1CC[C@H]2[C@@H]3CCC4=CC(=O)CC[C@]4(C)[C@H]3CC[C@]12C | neurosteroid | N.A | N.A | 0 | N.A | Ki(uM) | Inactive | N.A | N.A | 10.1016/S0028-3908(01)00071-5 | |
| Pregnenolone-succinate | 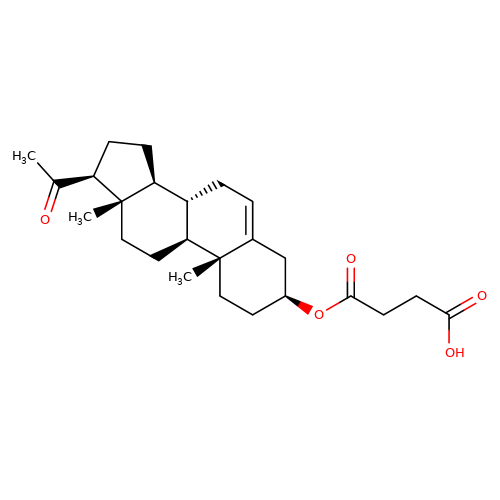 | CC(=O)[C@H]1CC[C@H]2[C@@H]3CC=C4C[C@@H](OC(=O)CCC(=O)O)CC[C@]4(C)[C@H]3CC[C@]12C | neurosteroid | (-)-neurosteroid | 5 | - | N.A | Ki(uM) | 9.8 | N.A | N.A | 10.1016/S0028-3908(01)00071-5 | |
| Pregnenolone-sulfate | 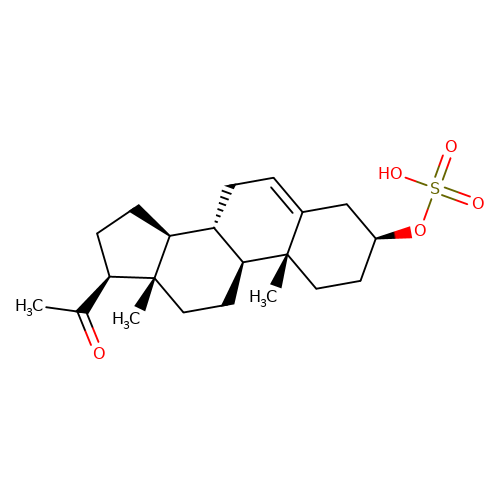 | CC(=O)[C@H]1CC[C@H]2[C@@H]3CC=C4C[C@@H](OS(=O)(=O)O)CC[C@]4(C)[C@H]3CC[C@]12C | neurosteroid | (-)-neurosteroid | 2 | - | N.A | Ki(uM) | 1.9 | N.A | N.A | 10.1016/S0028-3908(01)00071-5 | |
| THDOC | 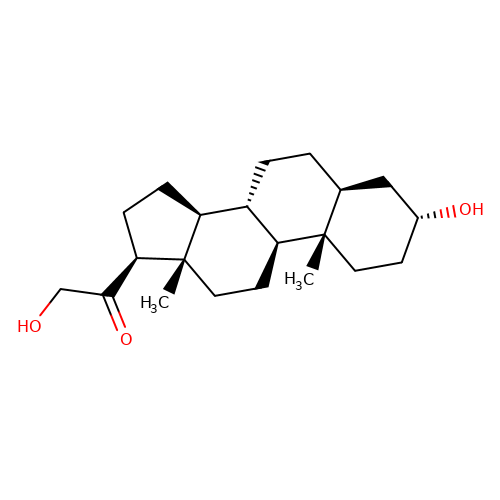 | C[C@]12CC[C@@H](O)C[C@@H]1CC[C@@H]1[C@@H]2CC[C@]2(C)[C@@H](C(=O)CO)CC[C@@H]12 | neurosteroid | (+)-neurosteroid | 2 | + | N.A | Maximum_effect(%) | 161 | N.A | N.A | 10.1113/JP272122 | |
| Dehydroepiandrosterone-sulfate | 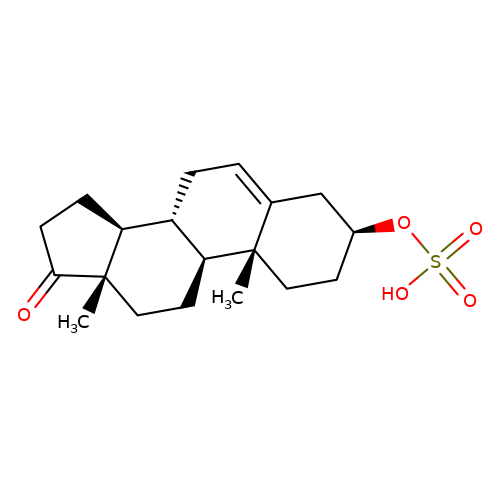 | C[C@]12CC[C@H]3[C@@H](CC=C4C[C@@H](OS(=O)(=O)O)CC[C@@]43C)[C@@H]1CCC2=O | neurosteroid | (-)-neurosteroid | 5 | - | N.A | Ki(uM) | 6.3 | N.A | N.A | 10.1016/S0028-3908(01)00071-5 | |
| Androsterone-sulfate | 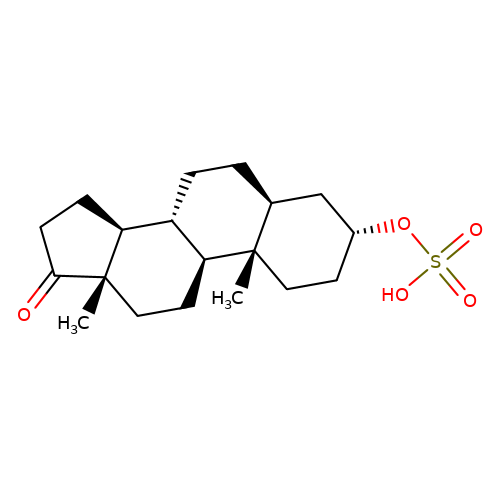 | C[C@]12CC[C@H]3[C@@H](CC[C@H]4C[C@H](OS(=O)(=O)O)CC[C@@]43C)[C@@H]1CCC2=O | neurosteroid | (-)-neurosteroid | 5 | - | N.A | Ki(uM) | 2.3 | N.A | N.A | 10.1016/S0028-3908(01)00071-5 | |
| androsterone | 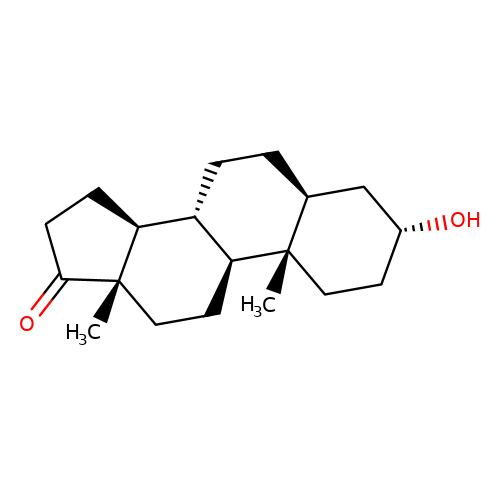 | C[C@]12CC[C@H]3[C@@H](CC[C@H]4C[C@H](O)CC[C@@]43C)[C@@H]1CCC2=O | neurosteroid | (+)-neurosteroid | 5 | + | N.A | Maximum_effect(%) | 317 | N.A | N.A | 10.1016/S0028-3908(01)00071-5 | |
| RU-5135 | 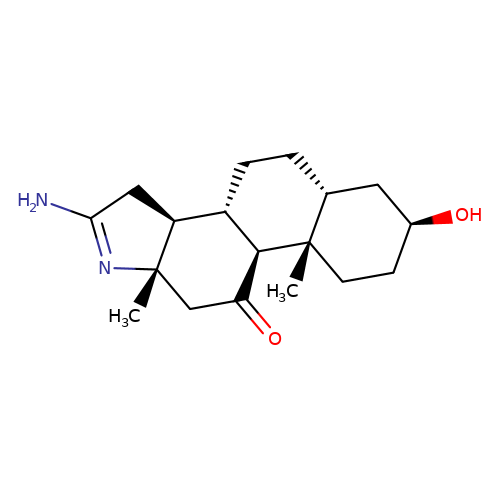 | C[C@]12CC(=O)[C@H]3[C@@H](CC[C@@H]4C[C@@H](O)CC[C@]34C)[C@@H]1CC(N)=N2 | neurosteroid | (+)-neurosteroid | 5 | - | - | IC50(uM) | 0.0531 | IC50(uM) | 0.0531 | 10.1111/j.1471-4159.1989.tb07256.x | |
| pregnanolone | 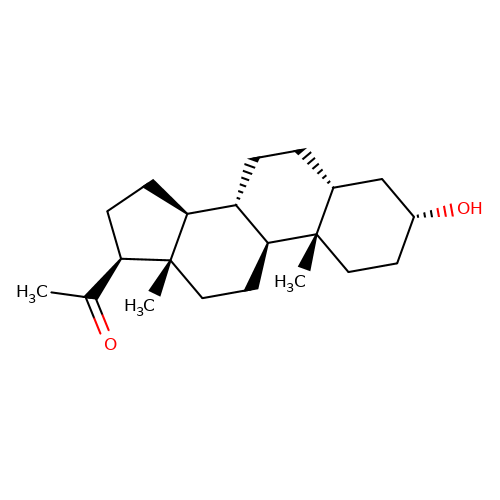 | CC(=O)[C@H]1CC[C@H]2[C@@H]3CC[C@@H]4C[C@H](O)CC[C@]4(C)[C@H]3CC[C@]12C | neurosteroid | (+)-neurosteroid | 2 | - | N.A | IC50(uM) | 1 | N.A | N.A | 10.1093/bja/aeh125 | |
| allopregnanolone | 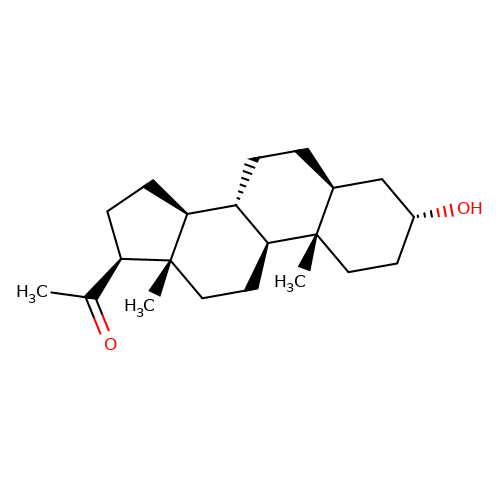 | CC(=O)[C@H]1CC[C@H]2[C@@H]3CC[C@H]4C[C@H](O)CC[C@]4(C)[C@H]3CC[C@]12C | neurosteroid | (+)-neurosteroid | 4 | + | N.A | EC50(uM) | 0.25 | N.A | N.A | 10.1093/bja/aeh125 | |
| sadek17-2 | 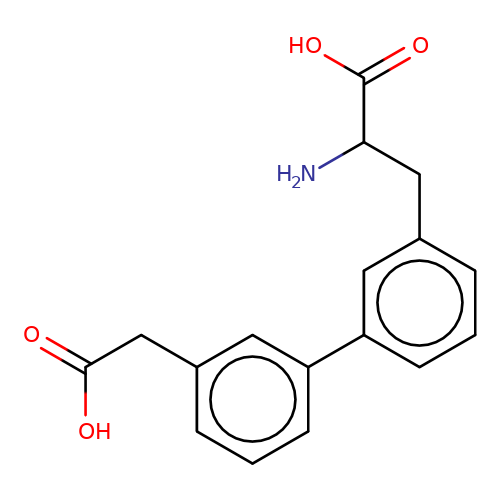 | NC(Cc1cccc(-c2cccc(CC(=O)O)c2)c1)C(=O)O | phenylalanine | N.A | N.A | - | N.A | Maximum_effect(%) | 5 | N.A | N.A | 10.1016/j.eplepsyres.2017.05.008 | |
| sadek17-6 | 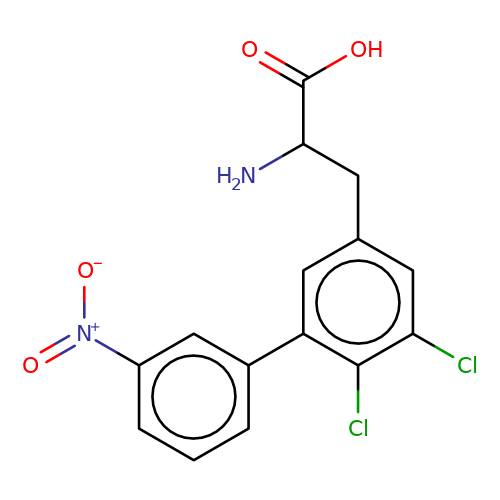 | NC(Cc1cc(Cl)c(Cl)c(-c2cccc([N+](=O)[O-])c2)c1)C(=O)O | phenylalanine | N.A | N.A | - | N.A | Maximum_effect(%) | 80 | N.A | N.A | 10.1016/j.eplepsyres.2017.05.008 | |
| sadek17-7 | 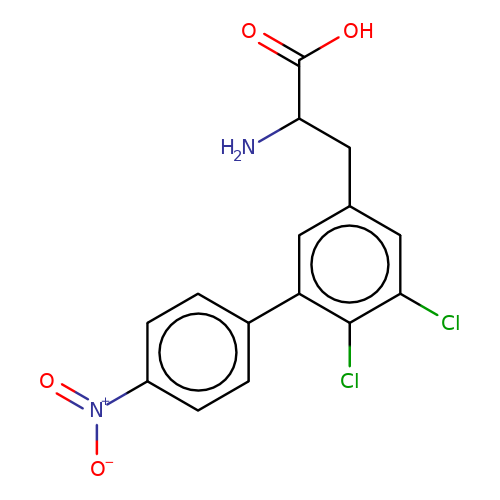 | NC(Cc1cc(Cl)c(Cl)c(-c2ccc([N+](=O)[O-])cc2)c1)C(=O)O | phenylalanine | N.A | N.A | - | N.A | Maximum_effect(%) | 90 | N.A | N.A | 10.1016/j.eplepsyres.2017.05.008 | |
| sadek17-1 | 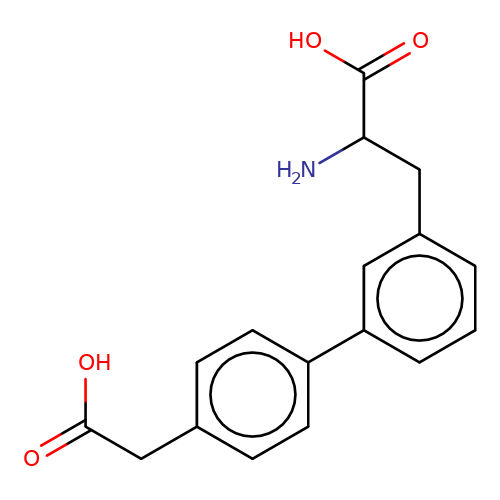 | NC(Cc1cccc(-c2ccc(CC(=O)O)cc2)c1)C(=O)O | phenylalanine | N.A | N.A | + | N.A | Maximum_effect(%) | 100 | N.A | N.A | 10.1016/j.eplepsyres.2017.05.008 | |
| sadek17-3 | 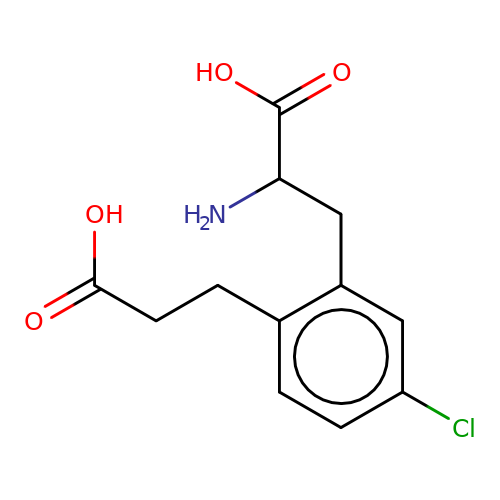 | NC(Cc1cc(Cl)ccc1CCC(=O)O)C(=O)O | phenylalanine | N.A | N.A | + | N.A | Maximum_effect(%) | 120 | N.A | N.A | 10.1016/j.eplepsyres.2017.05.008 | |
| sadek17-4 | 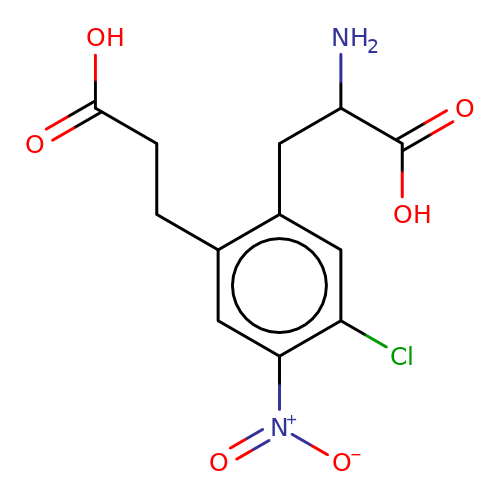 | NC(Cc1cc(Cl)c([N+](=O)[O-])cc1CCC(=O)O)C(=O)O | phenylalanine | N.A | N.A | + | N.A | Maximum_effect(%) | 110 | N.A | N.A | 10.1016/j.eplepsyres.2017.05.008 | |
| sadek17-5 | 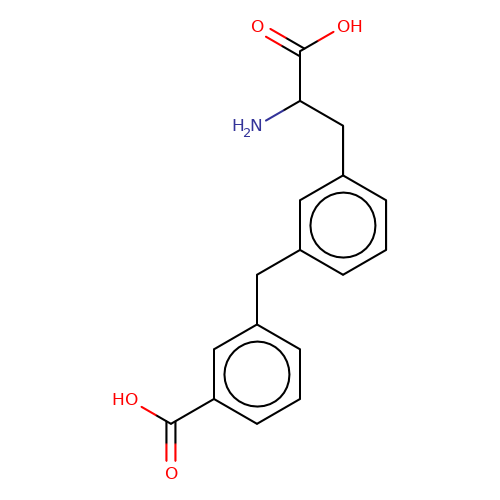 | NC(Cc1cccc(Cc2cccc(C(=O)O)c2)c1)C(=O)O | phenylalanine | N.A | N.A | + | N.A | Maximum_effect(%) | 160 | N.A | N.A | 10.1016/j.eplepsyres.2017.05.008 | |
| sadek17-8 | 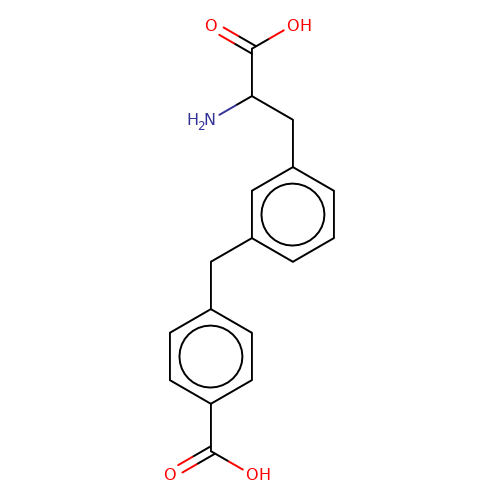 | NC(Cc1cccc(Cc2ccc(C(=O)O)cc2)c1)C(=O)O | phenylalanine | N.A | N.A | + | N.A | Maximum_effect(%) | 110 | N.A | N.A | 10.1016/j.eplepsyres.2017.05.008 | |
| fipronil | 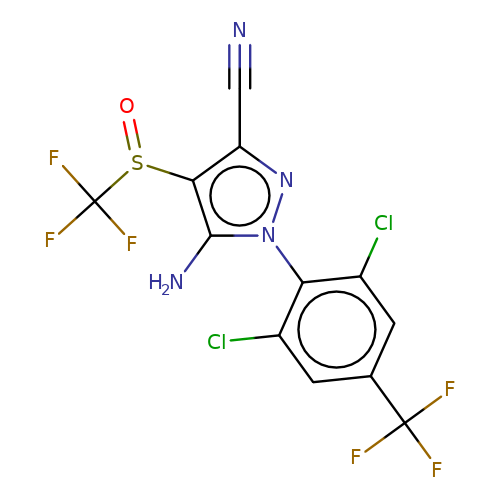 | N#Cc1nn(-c2c(Cl)cc(C(F)(F)F)cc2Cl)c(N)c1S(=O)C(F)(F)F | phenylpyrazole | pore | 3 | - | - | IC50(uM) | 1.96 | IC50(uM) | 0.35 | 10.1111/j.1476-5381.2011.01722.x | |
| lindane | 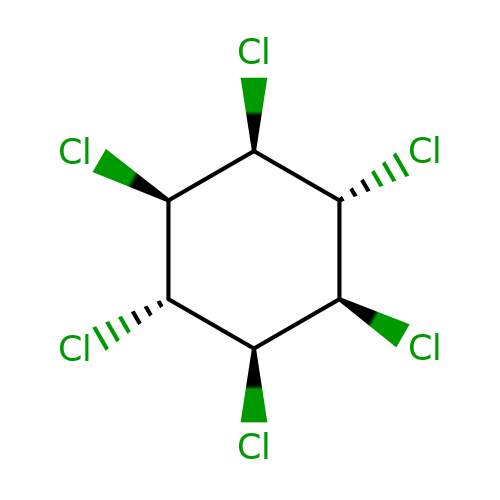 | Cl[C@H]1[C@H](Cl)[C@@H](Cl)[C@@H](Cl)[C@H](Cl)[C@H]1Cl | phenylpyrazole | pore | 3 | - | - | IC50(uM) | 0.9 | IC50(uM) | 0.25 | 10.1111/j.1476-5381.2011.01722.x | |
| picrotin | 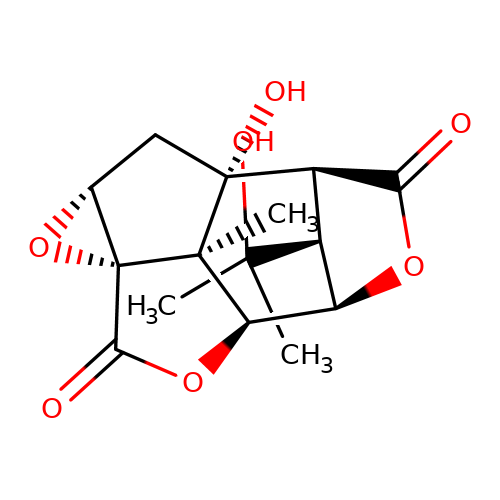 | CC(C)(O)[C@H]1[C@@H]2C(=O)O[C@H]1[C@H]1OC(=O)[C@@]34O[C@@H]3C[C@]2(O)[C@@]14C | picrotoxin | pore | 1 | - | - | IC50(uM) | 5.2 | IC50(uM) | 6 | 10.1111/j.1471-4159.2007.04850.x | |
| picrotoxinin | 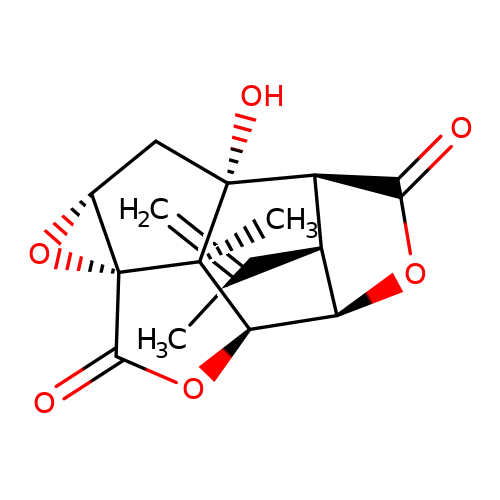 | C=C(C)[C@H]1[C@@H]2C(=O)O[C@H]1[C@H]1OC(=O)[C@@]34O[C@@H]3C[C@]2(O)[C@@]14C | picrotoxin | pore | 1 | - | - | IC50(uM) | 2.1 | IC50(uM) | 0.43 | 10.1111/j.1471-4159.2007.04850.x | |
| LT-01-25 | 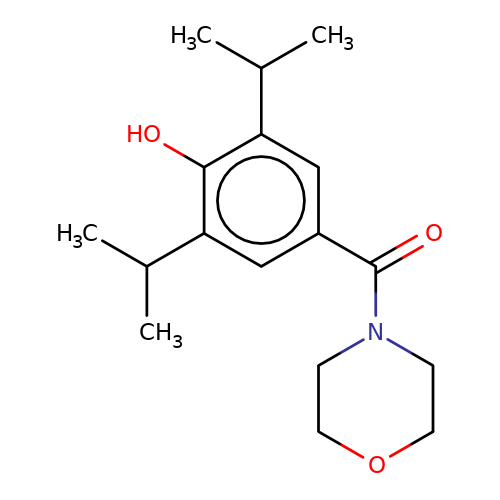 | CC(C)c1cc(C(=O)N2CCOCC2)cc(C(C)C)c1O | propofol | alcohol | 5 | + | N.A | EC50(uM) | 0.00005 | N.A | N.A | patent/US9676786B2 | |
| propofol | 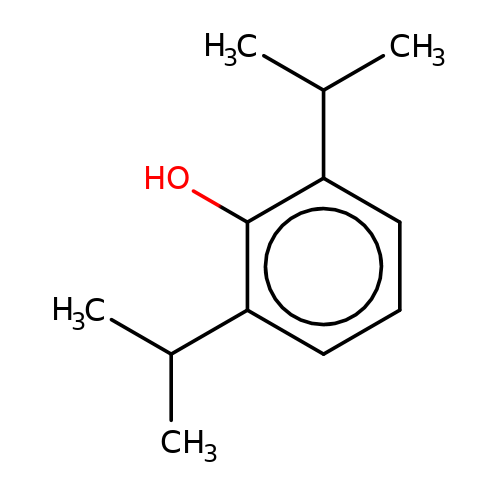 | CC(C)c1cccc(C(C)C)c1O | propofol | alcohol | 2 | + | N.A | EC50(uM)a1b | 12.5 | N.A | N.A | 10.1213/01.ANE.0000120083.10269.54 | |
| 2-6-DTBP | 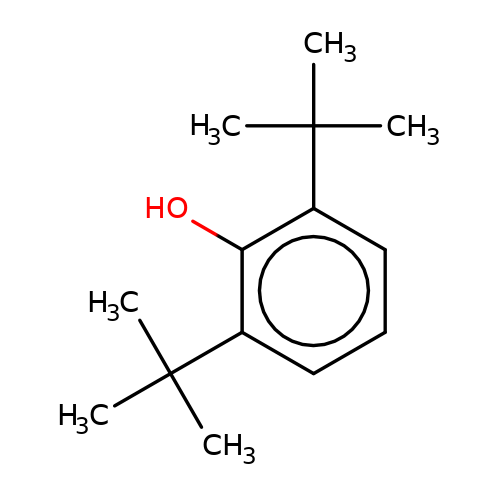 | CC(C)(C)c1cccc(C(C)(C)C)c1O | propofol | alcohol | 5 | + | + | EC50(uM)a1b | 9.4 | Mean_potention(%) | 309 | 10.1213/01.ANE.0000120083.10269.54 | |
| 4-chloropropofol | 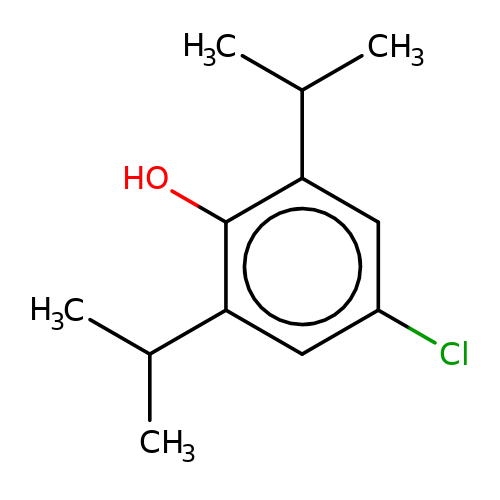 | CC(C)c1cc(Cl)cc(C(C)C)c1O | propofol | alcohol | 5 | + | + | EC50(uM) | 32 | EC50(uM) | 21 | 10.1111/bph.13566 | |
| 4-fluoropropofol | 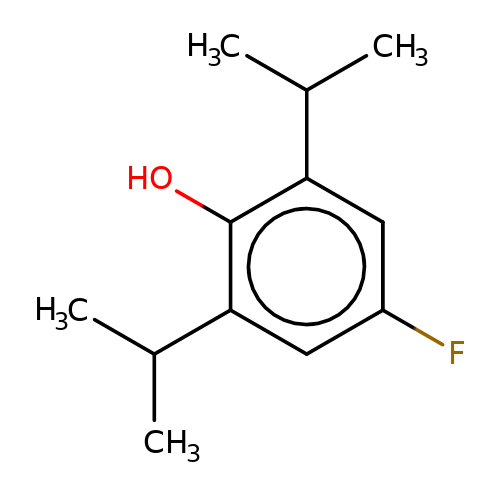 | CC(C)c1cc(F)cc(C(C)C)c1O | propofol | alcohol | 5 | + | + | EC50(uM) | 78 | EC50(uM) | 42 | 10.1111/bph.13566 | |
| 4-bromopropofol | 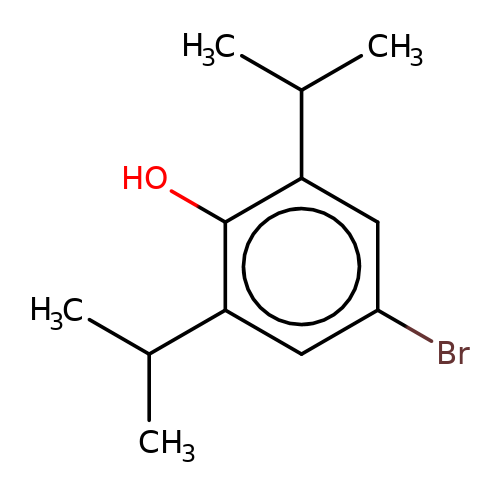 | CC(C)c1cc(Br)cc(C(C)C)c1O | propofol | alcohol | 5 | + | + | EC50(uM) | 42 | EC50(uM) | 11 | 10.1111/bph.13566 | |
| AM-1488 | 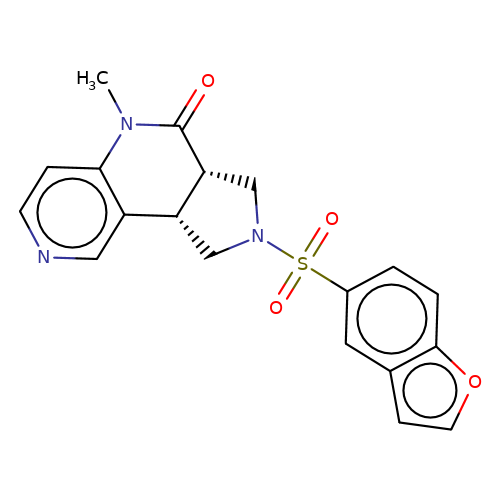 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4occc4c3)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | EC50(uM) | 0.33 | EC50(uM) | 0.45 | 10.1038/nsmb.3329 | |
| AM-3607 | 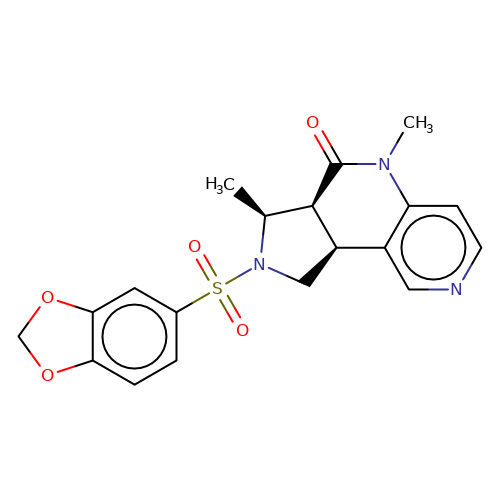 | C[C@H]1[C@H]2C(=O)N(C)c3ccncc3[C@H]2CN1S(=O)(=O)c1ccc2c(c1)OCO2 | sulfonamide | top_ECD | 1 | + | + | EC50(uM) | 0.03 | EC50(uM) | 0.05 | 10.1038/nsmb.3329 | |
| bregman-2 | 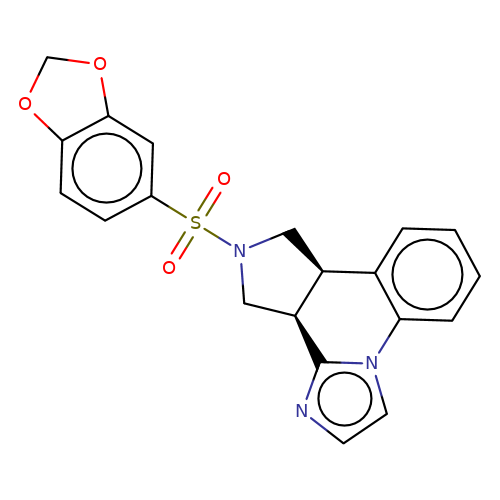 | O=S(=O)(c1ccc2c(c1)OCO2)N1C[C@H]2c3ccccc3-n3ccnc3[C@H]2C1 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 3.1 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-1 | 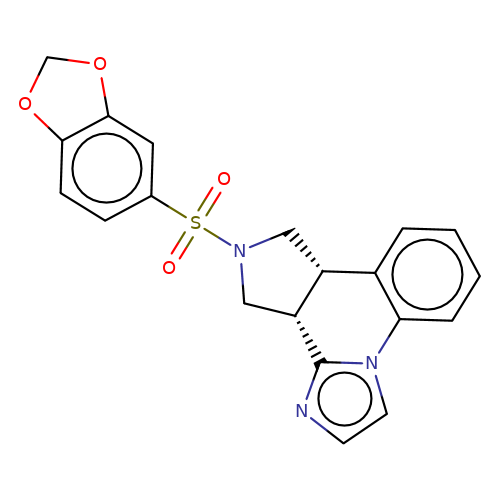 | O=S(=O)(c1ccc2c(c1)OCO2)N1C[C@@H]2c3ccccc3-n3ccnc3[C@@H]2C1 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 6.3 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-3 | 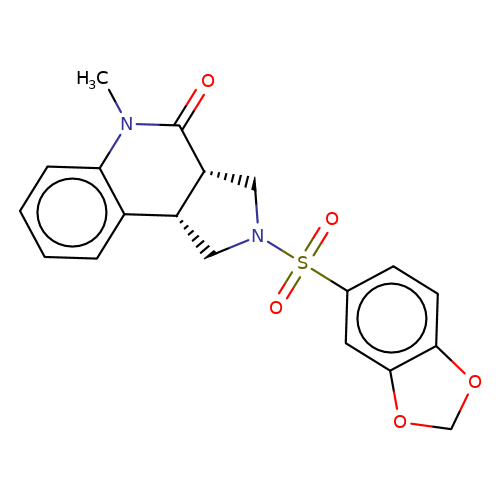 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@H]2c2ccccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 0.78 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-4 | 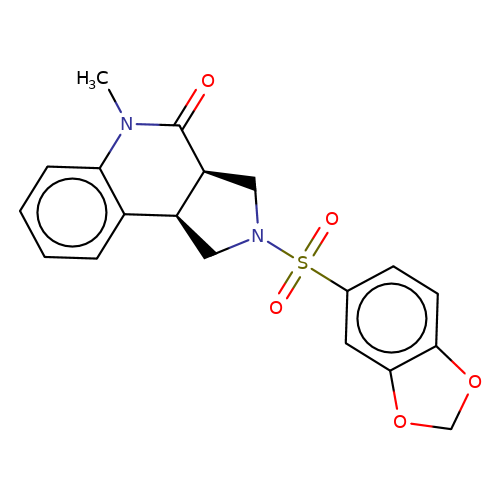 | CN1C(=O)[C@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@H]2c2ccccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 6.3 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-5 | 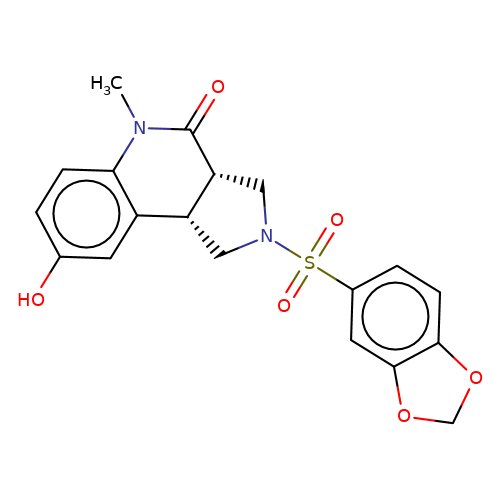 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@H]2c2cc(O)ccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 0.39 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-rac-6 | 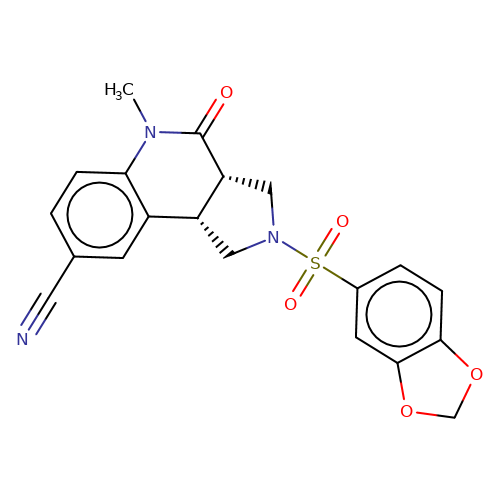 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@H]2c2cc(C#N)ccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 12.5 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-ent-6 | 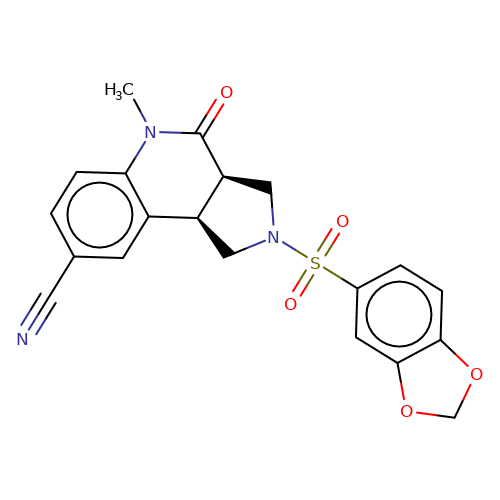 | CN1C(=O)[C@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@H]2c2cc(C#N)ccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 12.5 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-rac-7 | 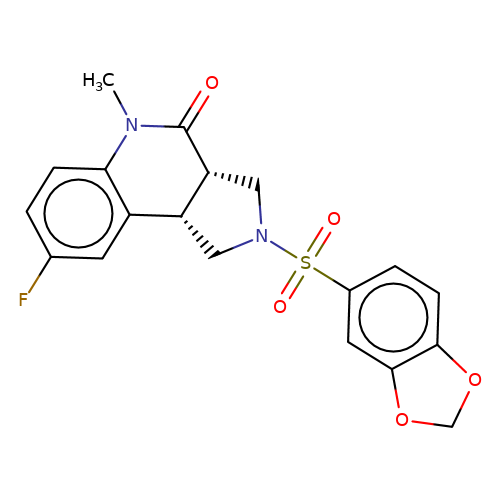 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@H]2c2cc(F)ccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 0.78 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-ent-7 | 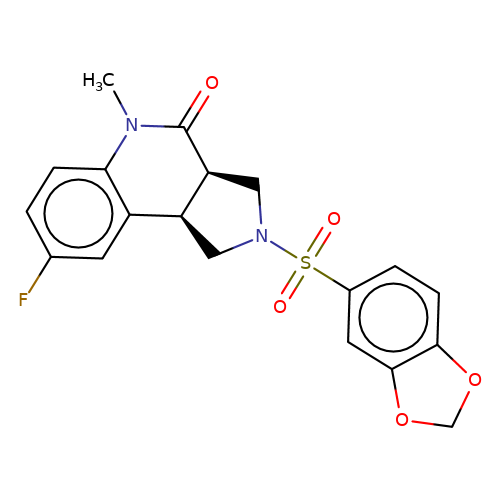 | CN1C(=O)[C@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@H]2c2cc(F)ccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 0.78 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-8 | 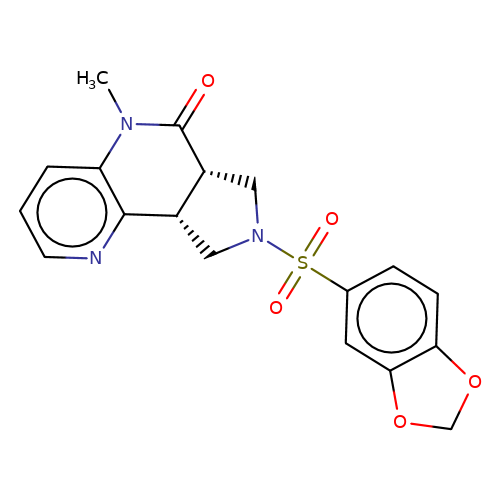 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@H]2c2ncccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 0.78 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-9 | 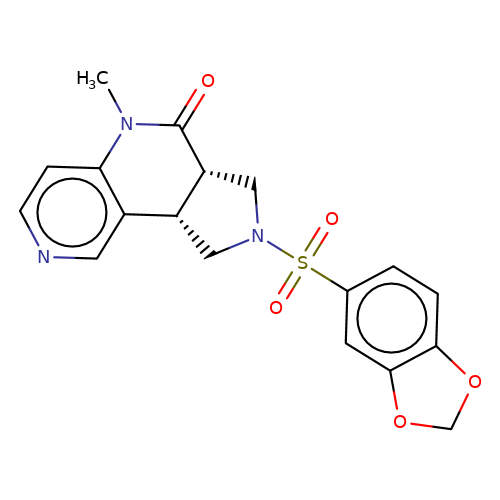 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 0.2 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-rac-10 | 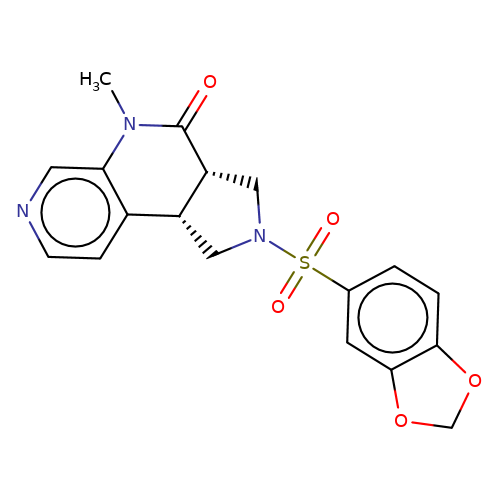 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@H]2c2ccncc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 6.3 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-ent-10 | 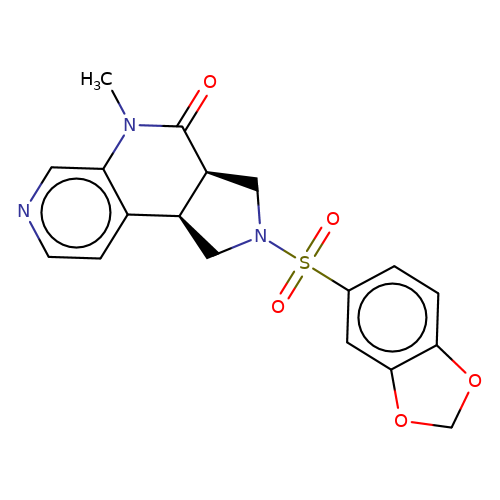 | CN1C(=O)[C@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@H]2c2ccncc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 6.3 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-rac-11 | 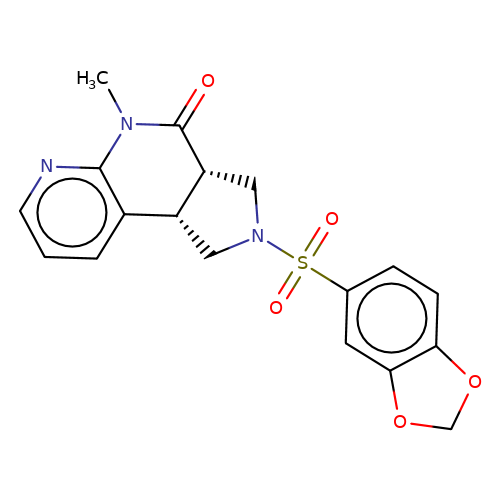 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@H]2c2cccnc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 3.1 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-ent-11 | 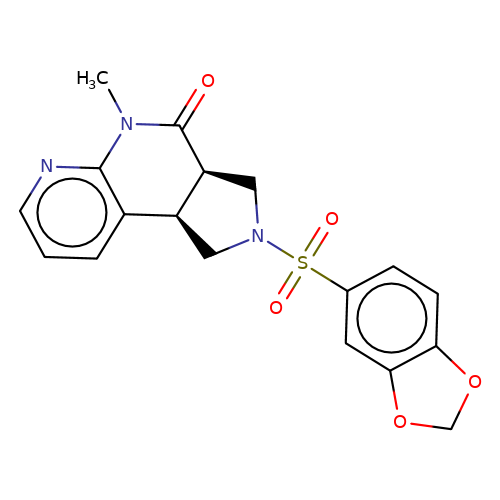 | CN1C(=O)[C@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@H]2c2cccnc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 3.1 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-12 | 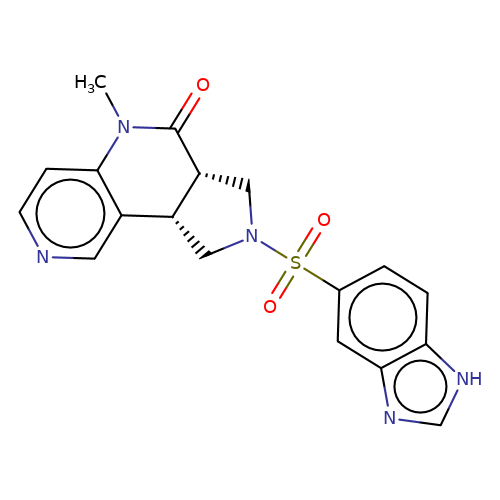 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4[nH]cnc4c3)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 3.1 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-13 | 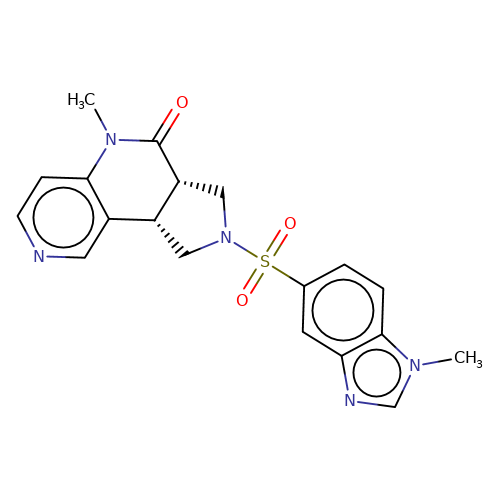 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)ncn4C)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 3.1 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-14 | 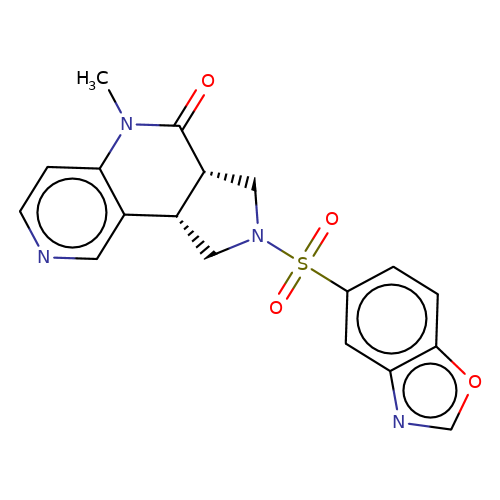 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4ocnc4c3)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 0.39 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-15 | 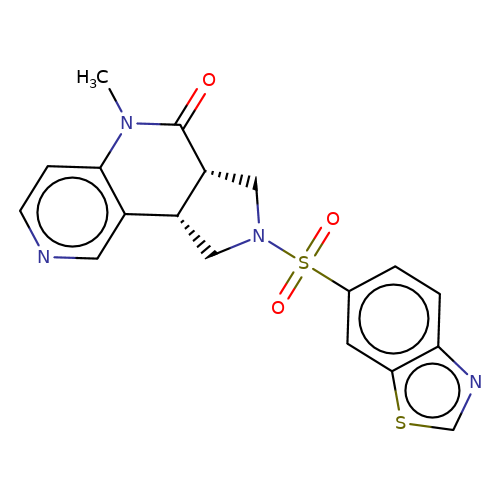 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4ncsc4c3)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 1.56 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-16 | 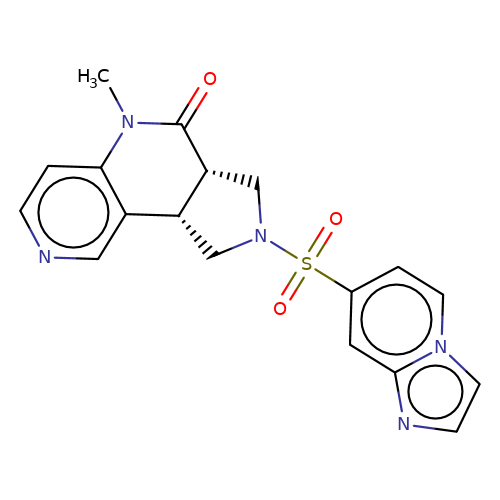 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccn4ccnc4c3)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 3.1 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-17 | 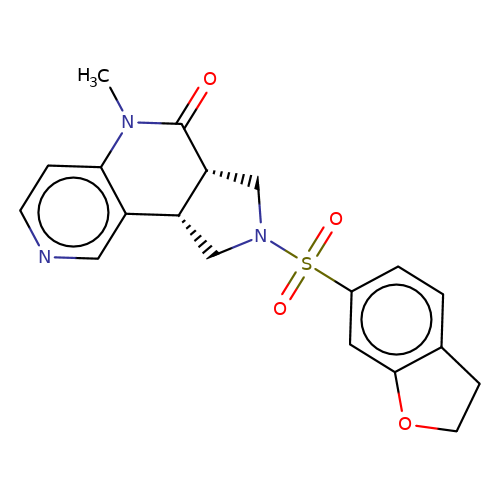 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)OCC4)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 1.56 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-18 | 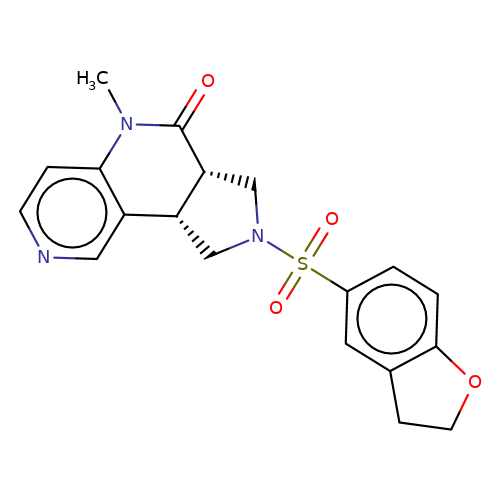 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)CCO4)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 3.1 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-19 | 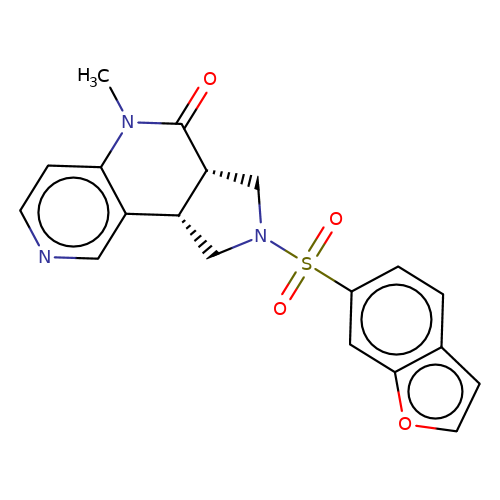 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4ccoc4c3)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 1.56 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-20 | 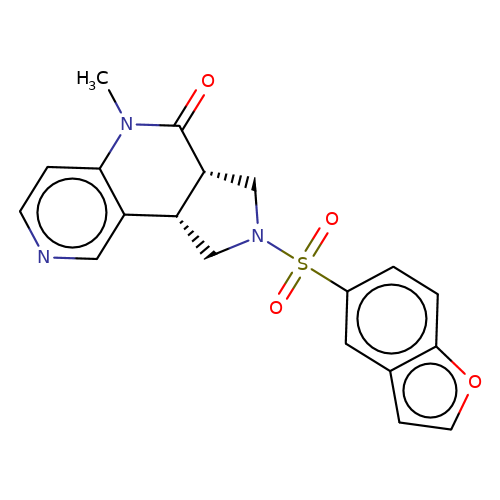 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4occc4c3)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 0.2 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-21 | 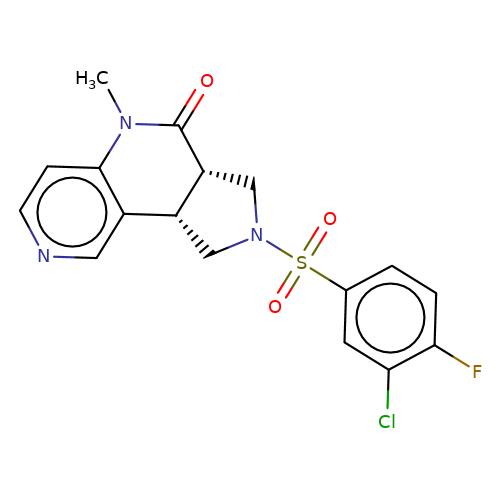 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc(F)c(Cl)c3)C[C@@H]2c2cnccc21 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 3.1 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-32 | 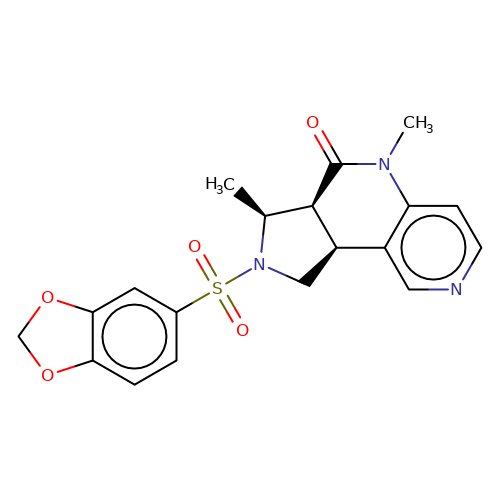 | C[C@H]1[C@H]2C(=O)N(C)c3ccncc3[C@H]2CN1S(=O)(=O)c1ccc2c(c1)OCO2 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 0.03 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-33 | 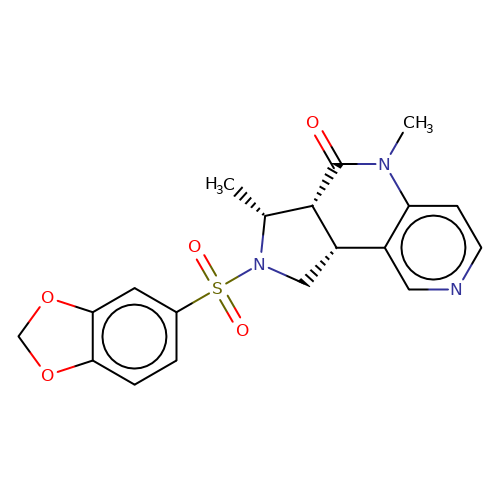 | C[C@@H]1[C@@H]2C(=O)N(C)c3ccncc3[C@@H]2CN1S(=O)(=O)c1ccc2c(c1)OCO2 | sulfonamide | top_ECD | 5 | + | + | N.A | N.A | EC50(uM)a3b | 3.1 | 10.1021/acs.jmedchem.6b01496 | |
| bregman-ent-9 | 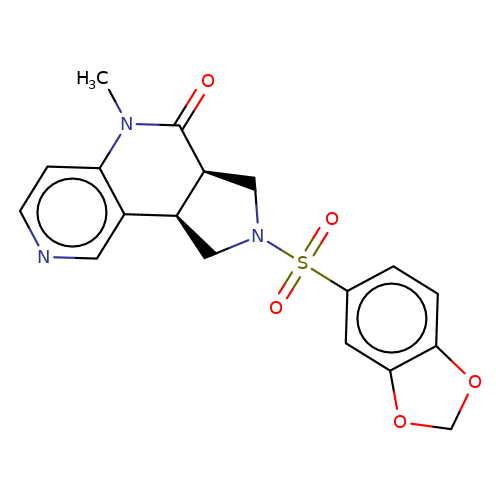 | CN1C(=O)[C@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@H]2c2cnccc21 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-ent-20 | 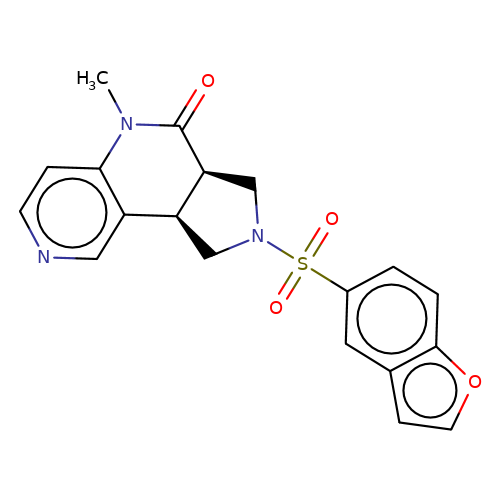 | CN1C(=O)[C@H]2CN(S(=O)(=O)c3ccc4occc4c3)C[C@H]2c2cnccc21 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-rac-22 | 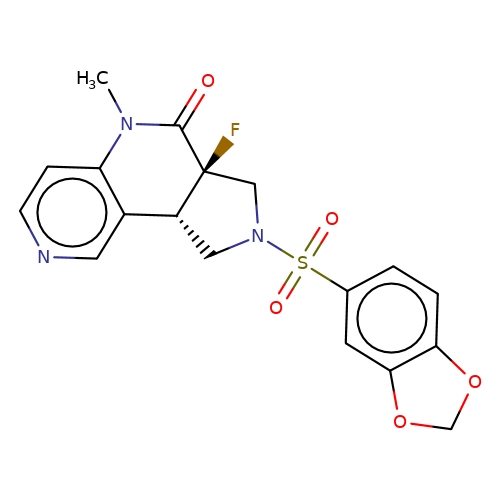 | CN1C(=O)[C@]2(F)CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@H]2c2cnccc21 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-ent-22 | 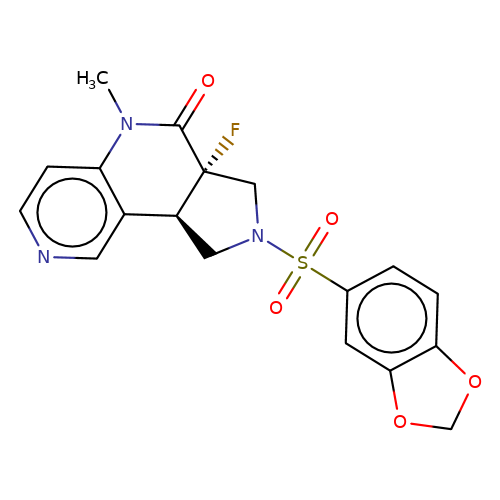 | CN1C(=O)[C@@]2(F)CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@H]2c2cnccc21 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-23 | 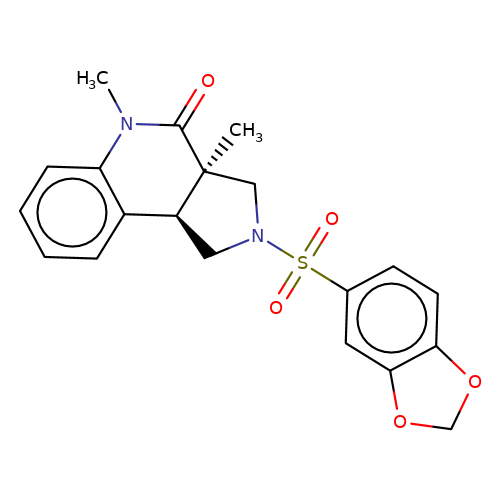 | CN1C(=O)[C@@]2(C)CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@H]2c2ccccc21 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-rac-24 | 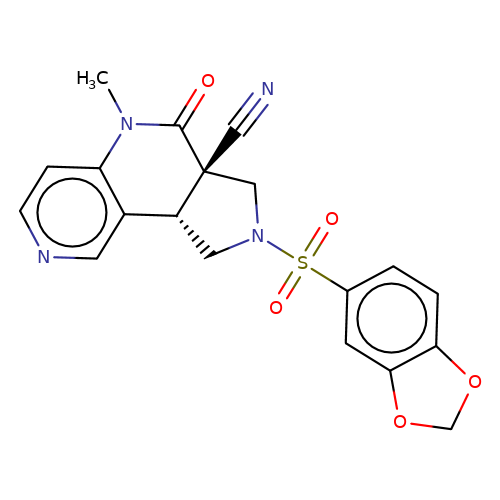 | CN1C(=O)[C@]2(C#N)CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@H]2c2cnccc21 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-ent-24 | 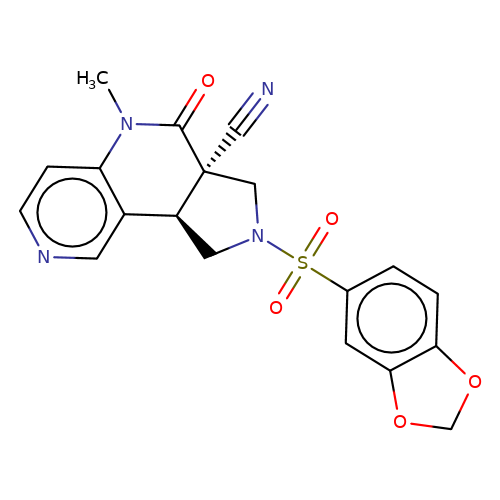 | CN1C(=O)[C@@]2(C#N)CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@H]2c2cnccc21 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-rac-25 | 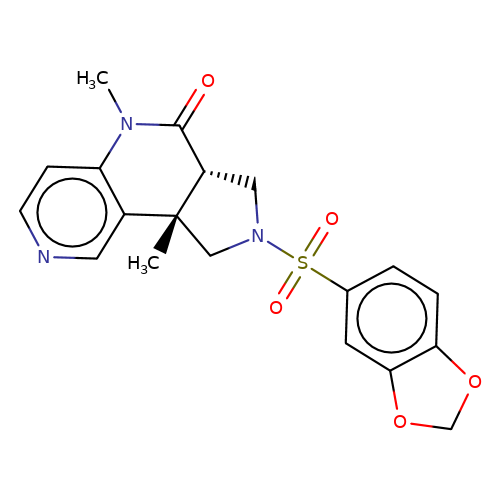 | CN1C(=O)[C@@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@]2(C)c2cnccc21 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-ent-25 | 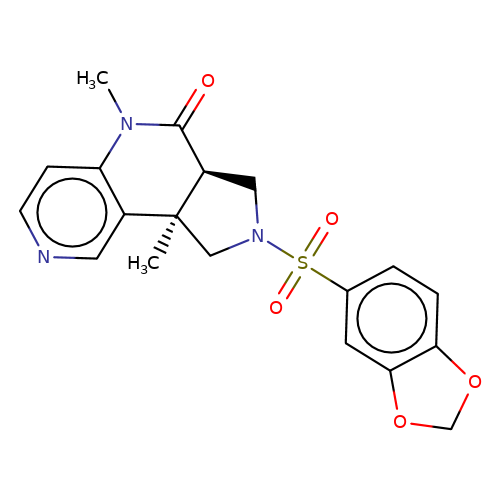 | CN1C(=O)[C@H]2CN(S(=O)(=O)c3ccc4c(c3)OCO4)C[C@@]2(C)c2cnccc21 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-26 | 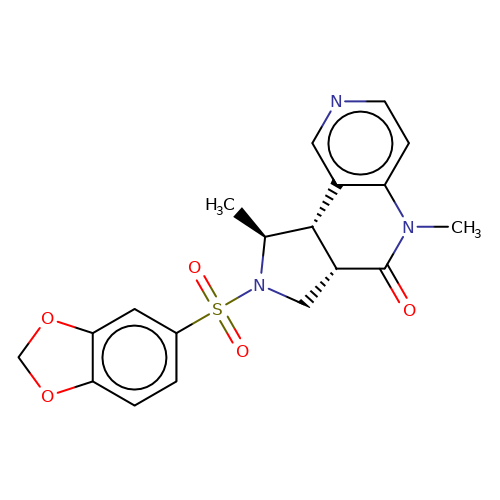 | C[C@H]1[C@@H]2c3cnccc3N(C)C(=O)[C@@H]2CN1S(=O)(=O)c1ccc2c(c1)OCO2 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-27 | 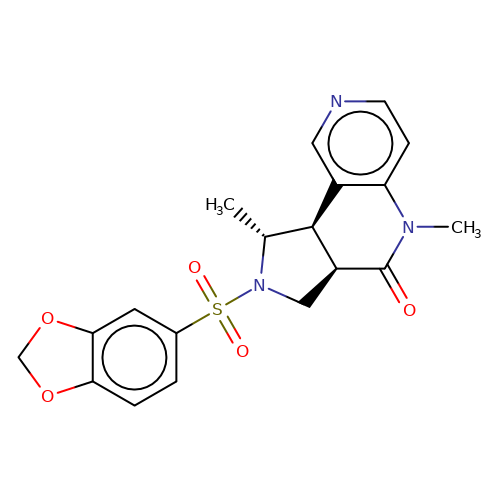 | C[C@@H]1[C@H]2c3cnccc3N(C)C(=O)[C@H]2CN1S(=O)(=O)c1ccc2c(c1)OCO2 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-28 | 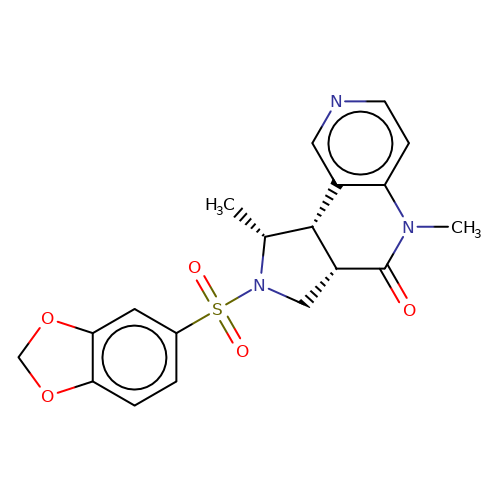 | C[C@@H]1[C@@H]2c3cnccc3N(C)C(=O)[C@@H]2CN1S(=O)(=O)c1ccc2c(c1)OCO2 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-29 | 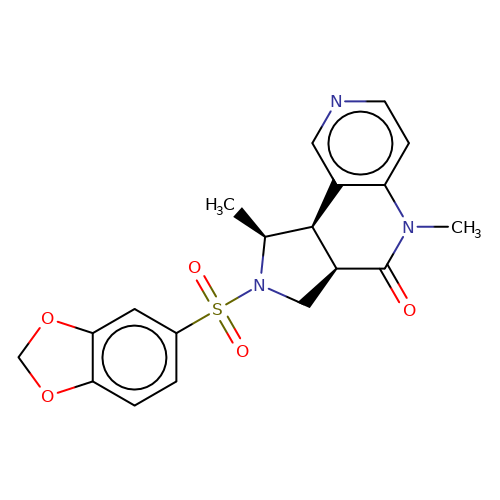 | C[C@H]1[C@H]2c3cnccc3N(C)C(=O)[C@H]2CN1S(=O)(=O)c1ccc2c(c1)OCO2 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-30 | 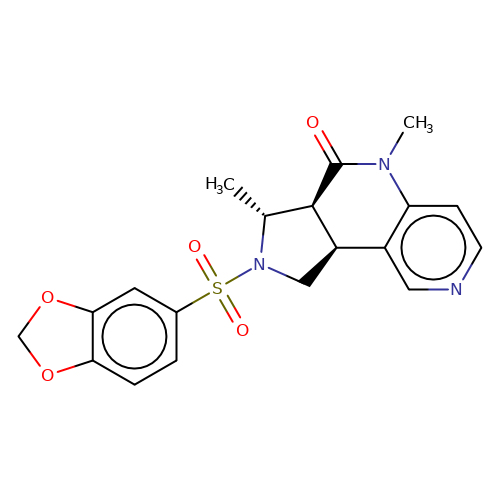 | C[C@@H]1[C@H]2C(=O)N(C)c3ccncc3[C@H]2CN1S(=O)(=O)c1ccc2c(c1)OCO2 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| bregman-31 | 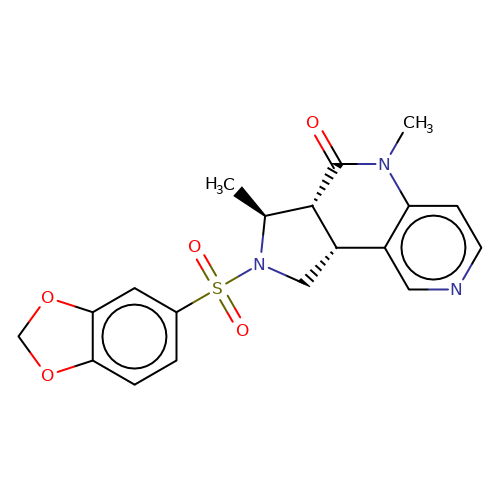 | C[C@H]1[C@@H]2C(=O)N(C)c3ccncc3[C@@H]2CN1S(=O)(=O)c1ccc2c(c1)OCO2 | sulfonamide | N.A | N.A | 0 | 0 | N.A | N.A | EC50(uM)a3b | Inactive | 10.1021/acs.jmedchem.6b01496 | |
| SB-203186 | 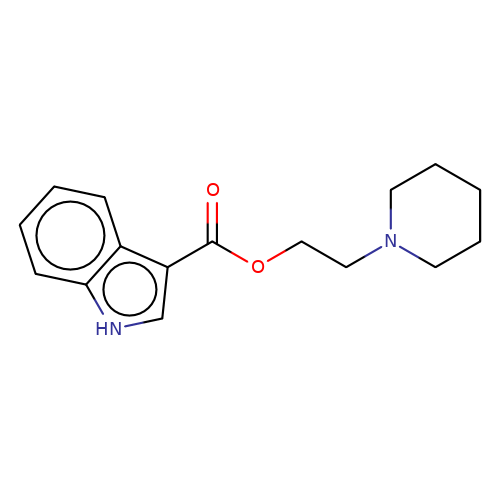 | O=C(OCCN1CCCCC1)c1c[nH]c2ccccc12 | tropeine | N.A | N.A | + | N.A | EC50(uM) | 0.01 | N.A | N.A | 10.1111/j.1471-4159.2006.04242.x | |
| MBN | 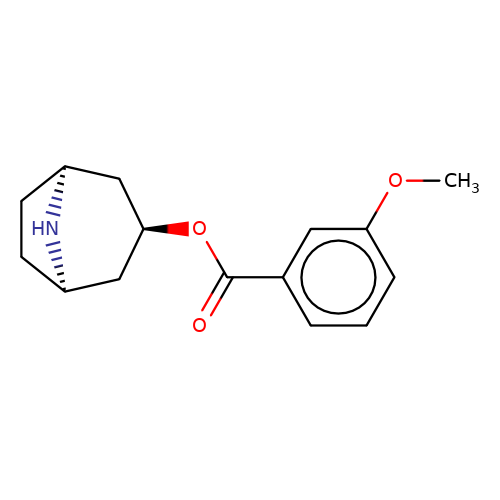 | COc1cccc(C(=O)O[C@@H]2C[C@@H]3CC[C@H](C2)N3)c1 | tropeine | N.A | N.A | + | N.A | IC50(uM) | 1.5 | N.A | N.A | 10.1111/j.1471-4159.2009.06083.x | |
| zatosetron | 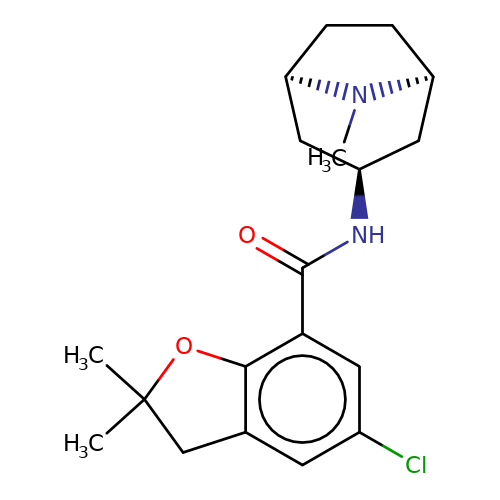 | CN1[C@H]2CC[C@@H]1C[C@H](NC(=O)c1cc(Cl)cc3c1OC(C)(C)C3)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.07 | Kd(uM) | 0.07 | 10.1016/S0028-3908(02)00213-7 | |
| bemesetron | 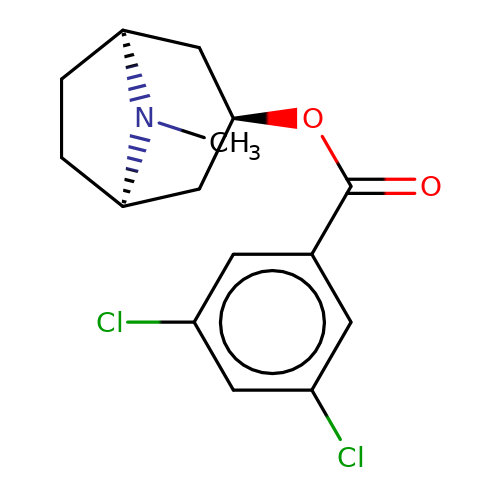 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)c1cc(Cl)cc(Cl)c1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.16 | Kd(uM) | 0.16 | 10.1016/S0028-3908(02)00213-7 | |
| tropisetron | 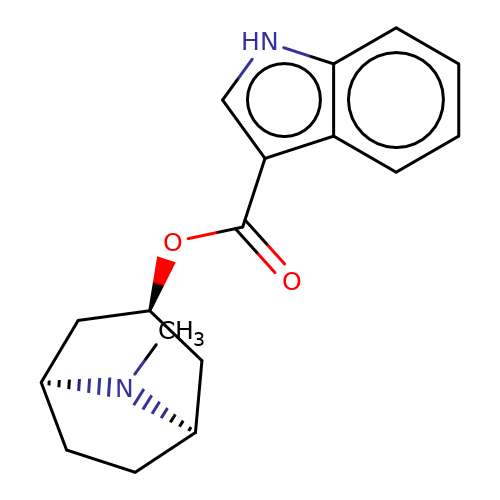 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)c1c[nH]c3ccccc13)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.09 | Kd(uM) | 0.09 | 10.1016/S0028-3908(02)00213-7 | |
| atropine | 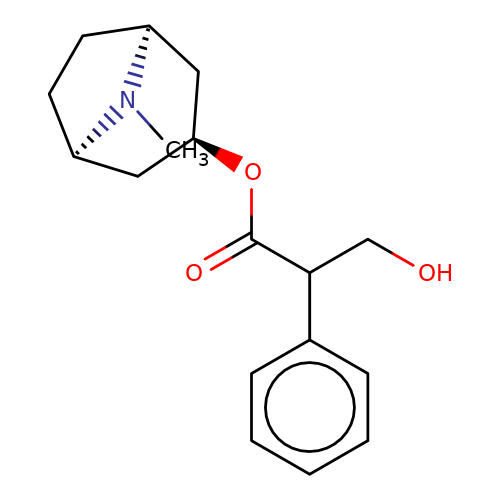 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)C(CO)c1ccccc1)C2 | tropeine | (LA)-tropeine | 5 | - | - | Kd(uM) | 1.6 | Kd(uM) | 1.6 | 10.1016/S0028-3908(02)00213-7 | |
| ondansetron | 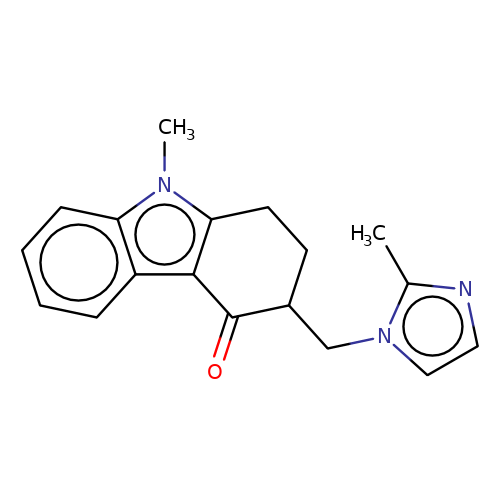 | Cc1nccn1CC1CCc2c(c3ccccc3n2C)C1=O | tropeine | (LA)-tropeine | 5 | - | - | Kd(uM) | 12.1 | Kd(uM) | 12.1 | 10.1016/S0028-3908(02)00213-7 | |
| granisetron | 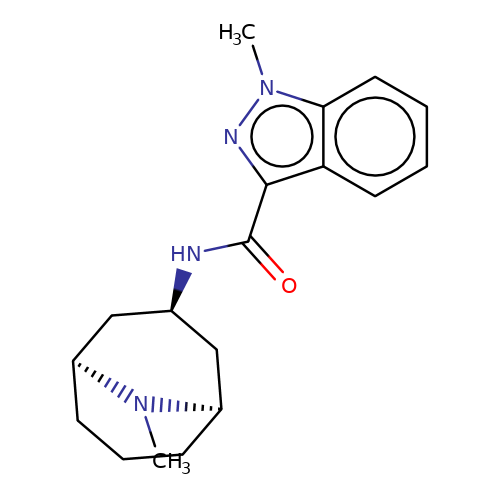 | CN1[C@H]2CCC[C@@H]1C[C@H](NC(=O)c1nn(C)c3ccccc13)C2 | tropeine | (LA)-tropeine | 2 | - | - | Kd(uM) | 12.5 | Kd(uM) | 12.5 | 10.1016/S0028-3908(02)00213-7 | |
| SR-57227A | 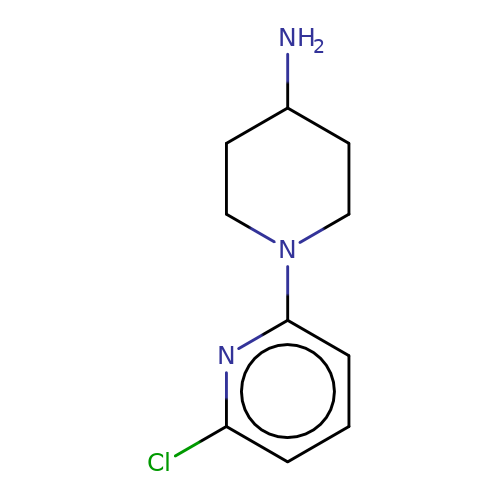 | NC1CCN(c2cccc(Cl)n2)CC1 | tropeine | (LA)-tropeine | 5 | - | - | Kd(uM) | 13.8 | Kd(uM) | 13.8 | 10.1016/S0028-3908(02)00213-7 | |
| M-chlorophenylbiguanide | 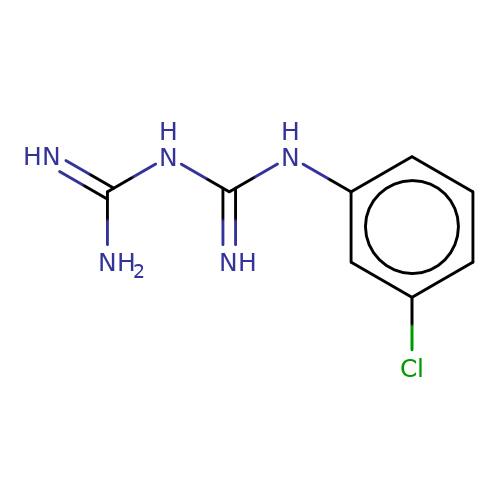 | N=C(N)NC(=N)Nc1cccc(Cl)c1 | tropeine | (LA)-tropeine | 5 | - | - | Kd(uM) | 25.4 | Kd(uM) | 25.4 | 10.1016/S0028-3908(02)00213-7 | |
| mianserin | 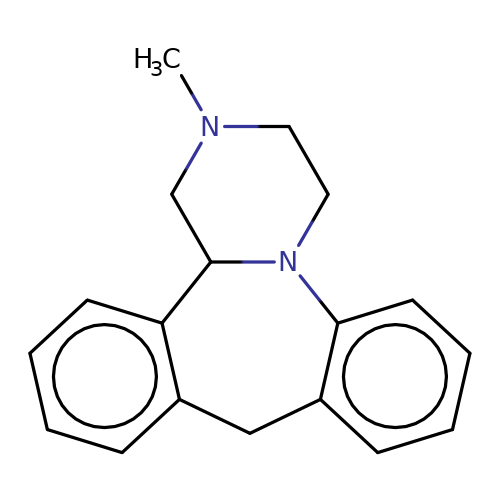 | CN1CCN2c3ccccc3Cc3ccccc3C2C1 | tropeine | (LA)-tropeine | 5 | - | - | Kd(uM) | 32.2 | Kd(uM) | 32.2 | 10.1016/S0028-3908(02)00213-7 | |
| scopolamine | 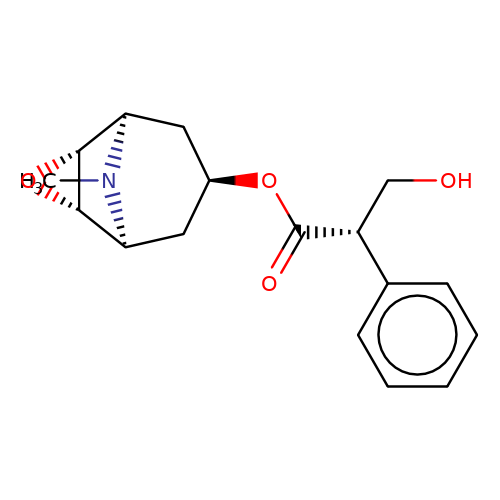 | CN1[C@H]2C[C@H](OC(=O)[C@H](CO)c3ccccc3)C[C@@H]1[C@H]1O[C@@H]21 | tropeine | (LA)-tropeine | 5 | - | - | Kd(uM) | 26 | Kd(uM) | 26 | 10.1016/S0028-3908(02)00213-7 | |
| metoclopramide | 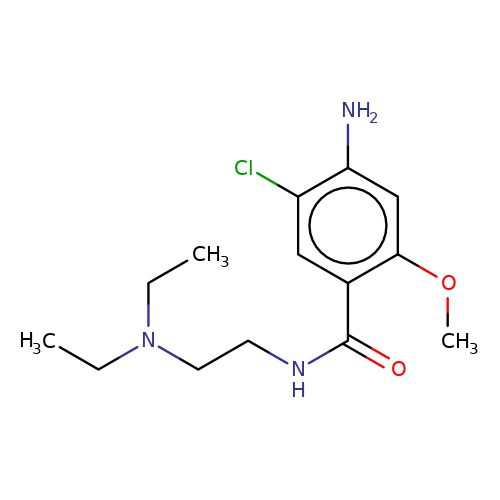 | CCN(CC)CCNC(=O)c1cc(Cl)c(N)cc1OC | tropeine | (LA)-tropeine | 5 | - | - | Kd(uM) | 229 | Kd(uM) | 229 | 10.1016/S0028-3908(02)00213-7 | |
| 3alpha-OH-tropane | 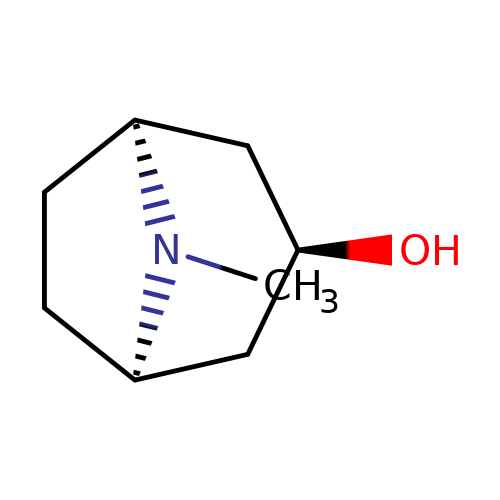 | CN1[C@H]2CC[C@@H]1C[C@H](O)C2 | tropeine | (LA)-tropeine | 5 | - | - | Kd(uM) | 290 | Kd(uM) | 290 | 10.1016/S0028-3908(02)00213-7 | |
| cocaine | 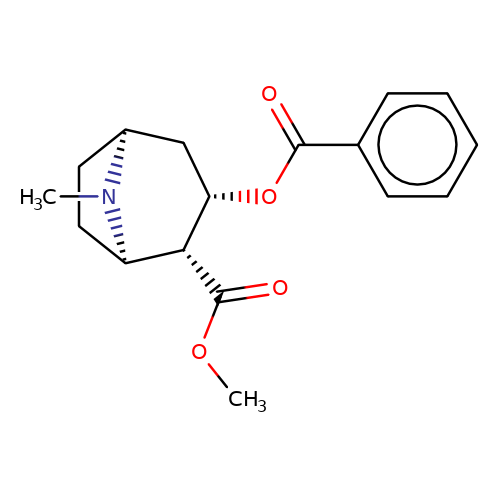 | COC(=O)[C@H]1[C@@H](OC(=O)c2ccccc2)C[C@@H]2CC[C@H]1N2C | tropeine | N.A | N.A | + | + | Kd(uM) | 208.65 | Kd(uM) | 208.65 | 10.1016/S0028-3908(02)00213-7 | |
| Maksay2004-2a | 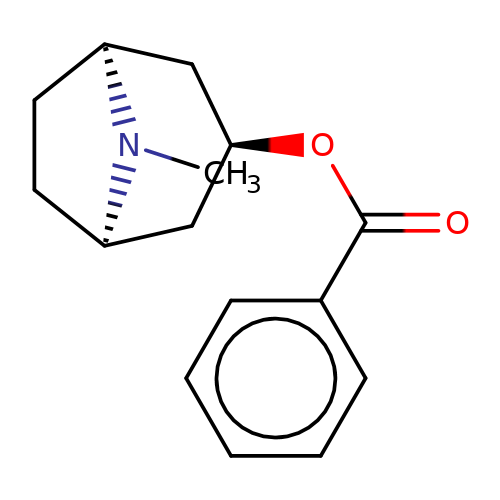 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)c1ccccc1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.2 | Kd(uM) | 0.2 | 10.1021/jm040814g | |
| Maksay2004-7 | 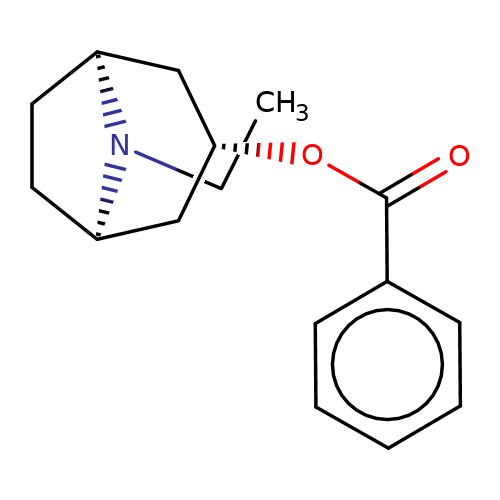 | CCN1[C@H]2CC[C@@H]1C[C@@H](OC(=O)c1ccccc1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.97 | Kd(uM) | 0.97 | 10.1021/jm040814g | |
| Maksay2004-2b | 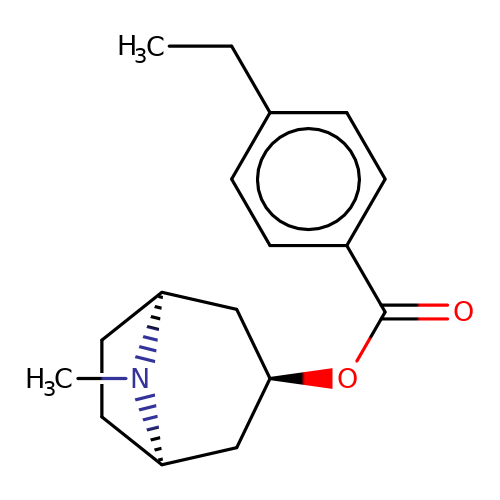 | CCc1ccc(C(=O)O[C@@H]2C[C@@H]3CC[C@H](C2)N3C)cc1 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.69 | Kd(uM) | 0.69 | 10.1021/jm040814g | |
| Maksay2004-2c | 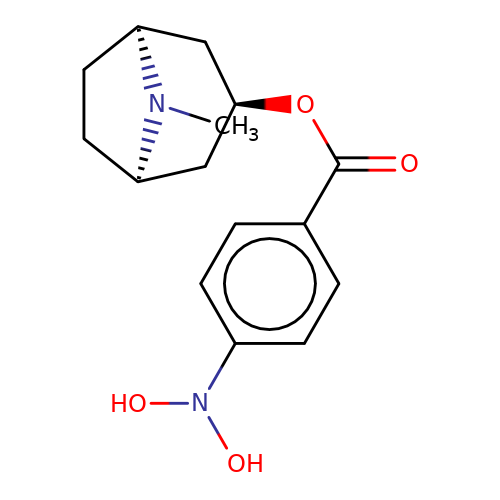 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)c1ccc(N(O)O)cc1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 1.02 | Kd(uM) | 1.02 | 10.1021/jm040814g | |
| Maksay2004-2d | 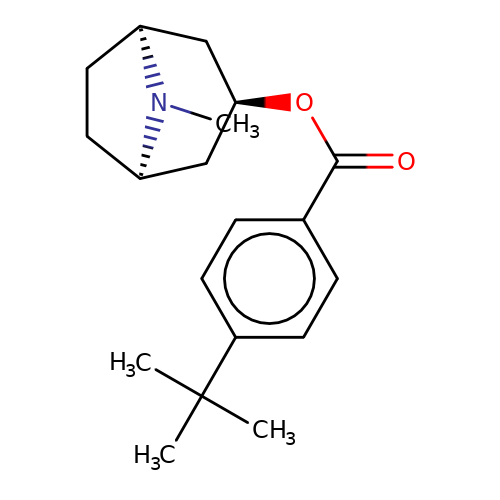 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)c1ccc(C(C)(C)C)cc1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 3.82 | Kd(uM) | 3.82 | 10.1021/jm040814g | |
| Maksay2004-2e | 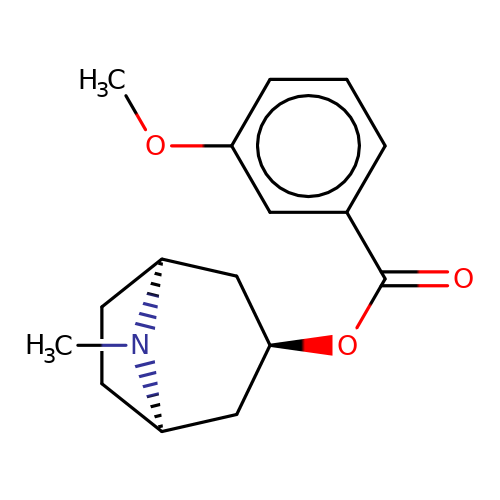 | COc1cccc(C(=O)O[C@@H]2C[C@@H]3CC[C@H](C2)N3C)c1 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.02 | Kd(uM) | 0.02 | 10.1021/jm040814g | |
| Maksay2004-2f | 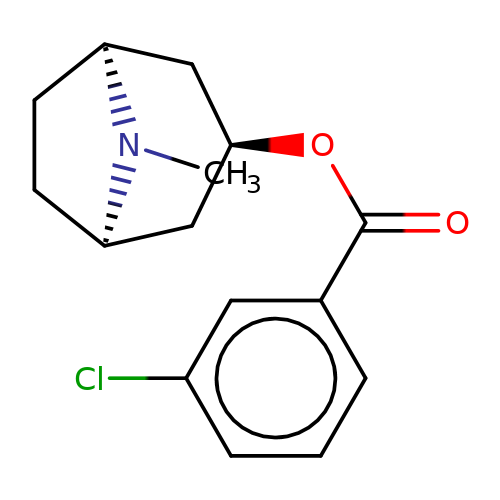 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)c1cccc(Cl)c1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.06 | Kd(uM) | 0.06 | 10.1021/jm040814g | |
| Maksay2004-2g | 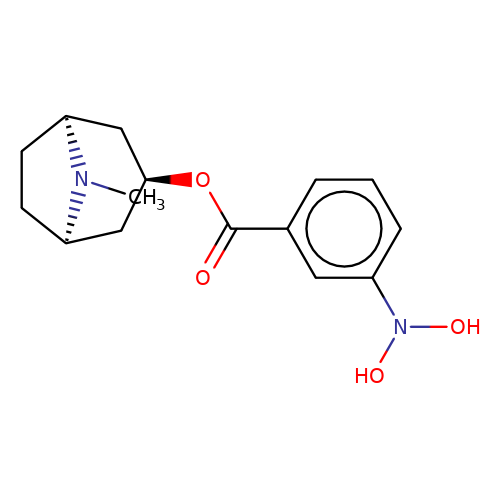 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)c1cccc(N(O)O)c1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.29 | Kd(uM) | 0.29 | 10.1021/jm040814g | |
| Maksay2004-2h | 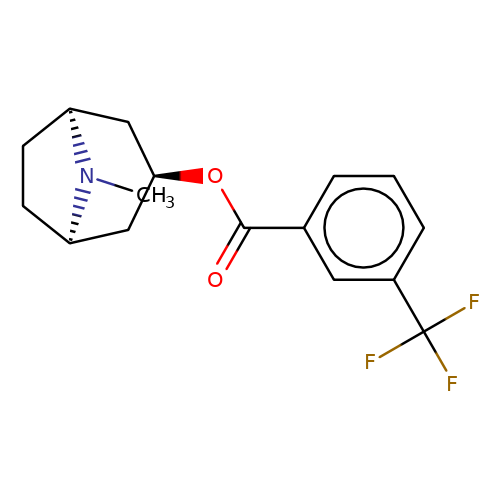 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)c1cccc(C(F)(F)F)c1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.18 | Kd(uM) | 0.18 | 10.1021/jm040814g | |
| Maksay2004-2i | 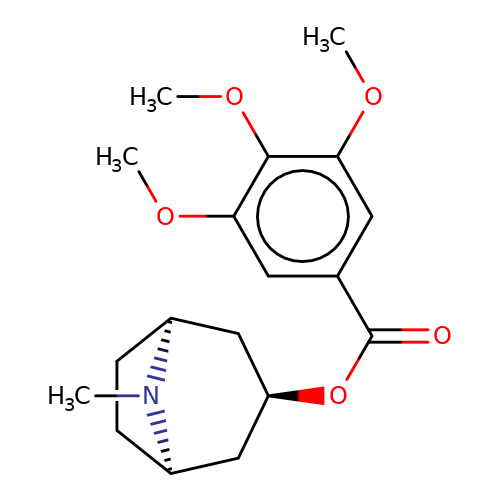 | COc1cc(C(=O)O[C@@H]2C[C@@H]3CC[C@H](C2)N3C)cc(OC)c1OC | tropeine | N.A | N.A | + | + | Kd(uM) | 0.1 | Kd(uM) | 0.1 | 10.1021/jm040814g | |
| Maksay2004-2j | 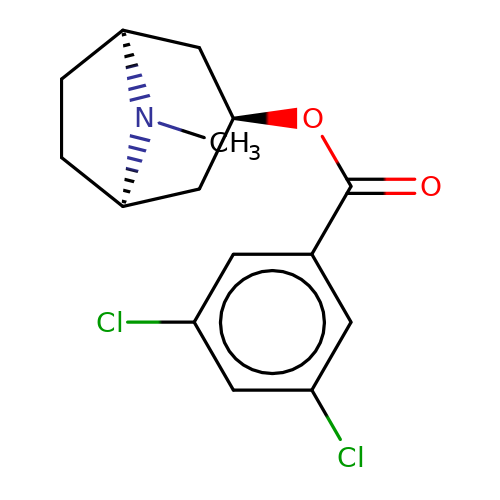 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)c1cc(Cl)cc(Cl)c1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.06 | Kd(uM) | 0.06 | 10.1021/jm040814g | |
| Maksay2004-8 | 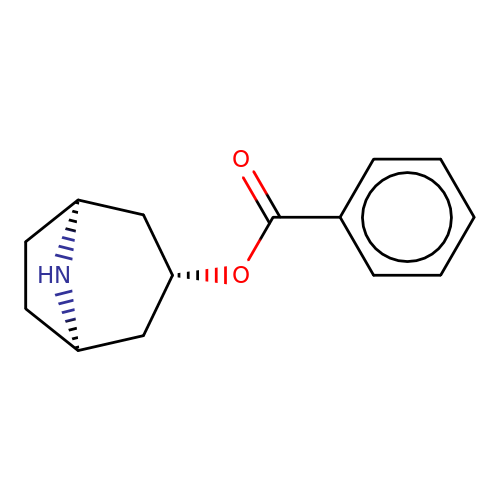 | O=C(O[C@H]1C[C@@H]2CC[C@H](C1)N2)c1ccccc1 | tropeine | (LA)-tropeine | 5 | - | - | Kd(uM) | 1.1 | Kd(uM) | 1.1 | 10.1021/jm040814g | |
| Maksay2004-4a | 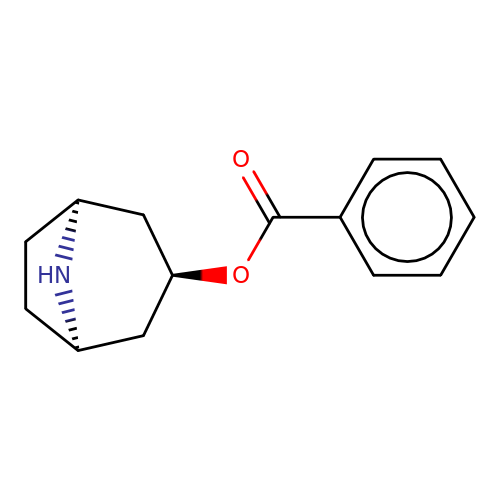 | O=C(O[C@@H]1C[C@@H]2CC[C@H](C1)N2)c1ccccc1 | tropeine | N.A | N.A | + | + | Kd(uM) | Inactive | Kd(uM) | Inactive | 10.1021/jm040814g | |
| Maksay2004-4b | 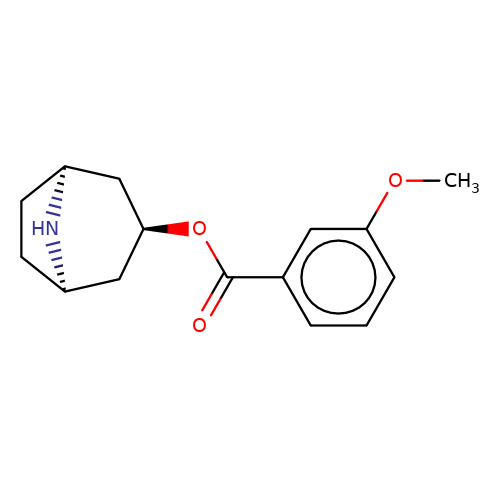 | COc1cccc(C(=O)O[C@@H]2C[C@@H]3CC[C@H](C2)N3)c1 | tropeine | N.A | N.A | + | + | Kd(uM) | Inactive | Kd(uM) | Inactive | 10.1021/jm040814g | |
| Maksay2004-4c | 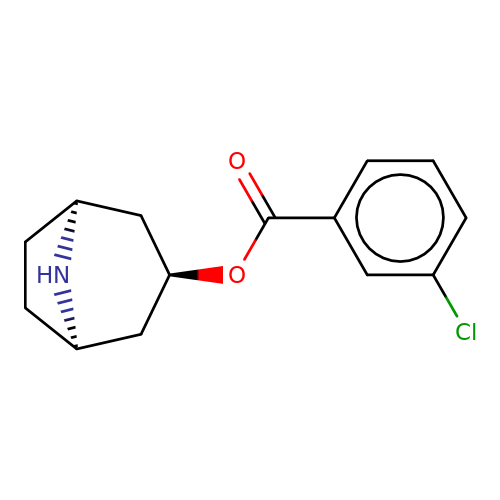 | O=C(O[C@@H]1C[C@@H]2CC[C@H](C1)N2)c1cccc(Cl)c1 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.01 | Kd(uM) | 0.01 | 10.1021/jm040814g | |
| Maksay2004-4e | 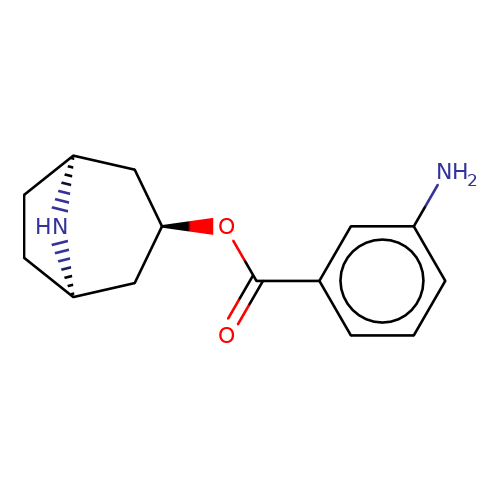 | Nc1cccc(C(=O)O[C@@H]2C[C@@H]3CC[C@H](C2)N3)c1 | tropeine | (LA)-tropeine | 5 | - | - | Kd(uM) | 0.06 | Kd(uM) | 0.06 | 10.1021/jm040814g | |
| Maksay2004-2k | 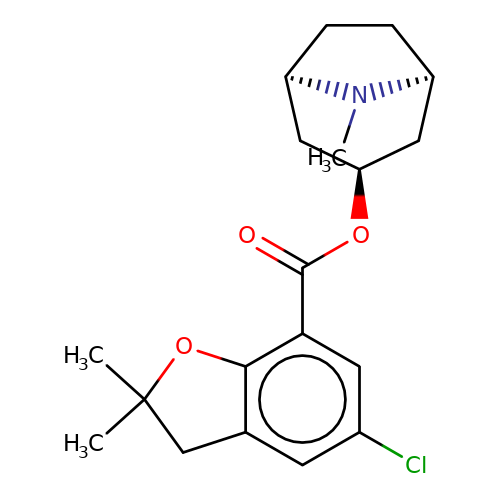 | CN1[C@H]2CC[C@@H]1C[C@H](OC(=O)c1cc(Cl)cc3c1OC(C)(C)C3)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.01 | Kd(uM) | 0.01 | 10.1021/jm040814g | |
| Maksay2004-4d | 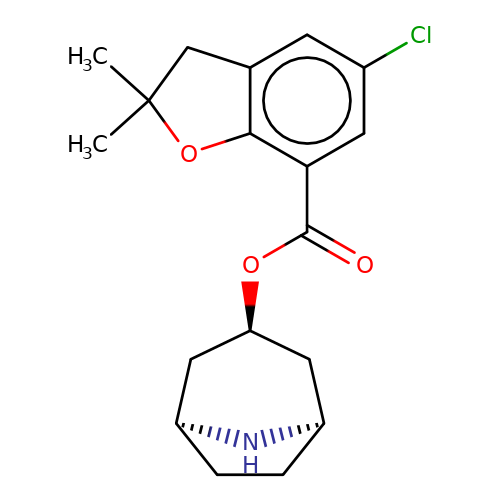 | CC1(C)Cc2cc(Cl)cc(C(=O)O[C@@H]3C[C@@H]4CC[C@H](C3)N4)c2O1 | tropeine | N.A | N.A | + | + | Kd(uM) | Inactive | Kd(uM) | Inactive | 10.1021/jm040814g | |
| Maksay2004-9 | 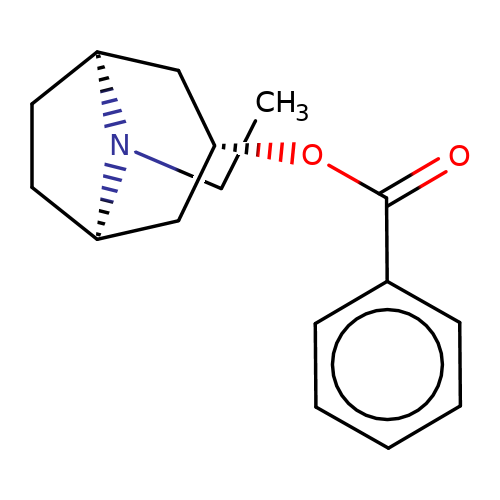 | CCN1[C@H]2CC[C@@H]1C[C@@H](OC(=O)c1ccccc1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 0.4 | Kd(uM) | 0.4 | 10.1021/jm040814g | |
| Maksay2004-10 | 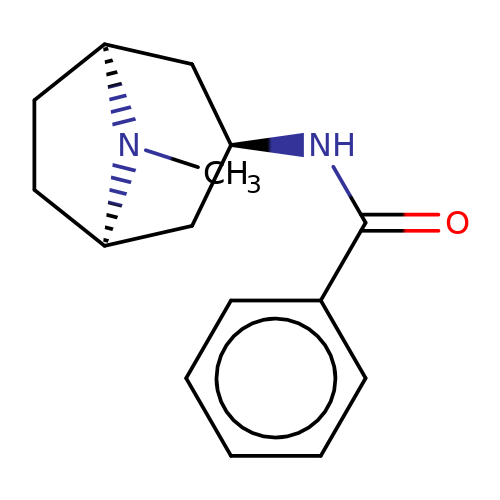 | CN1[C@H]2CC[C@@H]1C[C@H](NC(=O)c1ccccc1)C2 | tropeine | N.A | N.A | + | + | Kd(uM) | 3.52 | Kd(uM) | 3.52 | 10.1021/jm040814g |
Acknowledgments:


